Summary: Researchers have invented a probe that lights up when it binds to a misfolded amyloid peptide.
Source: Rice University.
A probe invented at Rice University that lights up when it binds to a misfolded amyloid beta peptide — the kind suspected of causing Alzheimer’s disease — has identified a specific binding site on the protein that could facilitate better drugs to treat the disease.
Even better, the lab has discovered that when the metallic probe is illuminated, it catalyzes oxidation of the protein in a way they believe might keep it from aggregating in the brains of patients.
The study done on long amyloid fibrils backs up computer simulations by colleagues at the University of Miami that predicted the photoluminescent metal complex would attach itself to the amyloid peptide near a hydrophobic (water-avoiding) cleft that appears on the surface of the fibril aggregate. That cleft presents a new target for drugs.
Finding the site was relatively simple once the lab of Rice chemist Angel Martí used its rhenium-based complexes to target fibrils. The light-switching complex glows when hit with ultraviolet light, but when it binds to the fibril it becomes more than 100 times brighter and causes oxidation of the amyloid peptide.
“It’s like walking on the beach,” Marti said. “You can see that someone was there before you by looking at footprints in the sand. While we cannot see the rhenium complex, we can find the oxidation (footprint) it produces on the amyloid peptide.
“That oxidation only happens right next to the place where it binds,” he said. “The real importance of this research is that allows us to see with a high degree of certainty where molecules can interact with amyloid beta fibrils.”
The study appears in the journal Chem.
“We believe this hydrophobic cleft is a general binding site (on amyloid beta) for molecules,” Martí said. “This is important because amyloid beta aggregation has been associated with the onset of Alzheimer’s disease. We know that fibrillar insoluble amyloid beta is toxic to cell cultures. Soluble amyloid oligomers that are made of several misfolded units of amyloid beta are also toxic to cells, probably even more than fibrillar.
“There’s an interest in finding medications that will quench the deleterious effects of amyloid beta aggregates,” he said. “But to create drugs for these, we first need to know how drugs or molecules in general can bind and interact with these fibrils, and this was not well-known. Now we have a better idea of what the molecule needs to interact with these fibrils.”
When amyloid peptides fold properly, they hide their hydrophobic residues while exposing their hydrophilic (water-attracting) residues to water. That makes the proteins soluble, Martí said. But when amyloid beta misfolds, it leaves two hydrophobic residues, known as Valine 18 and Phenylalanine 20, exposed to create the hydrophobic cleft.
“It’s perfect, because then molecules with hydrophobic domains are driven to bind there,” Martí said. “They are compatible with this hydrophobic cleft and associate with the fibril, forming a strong interaction.”
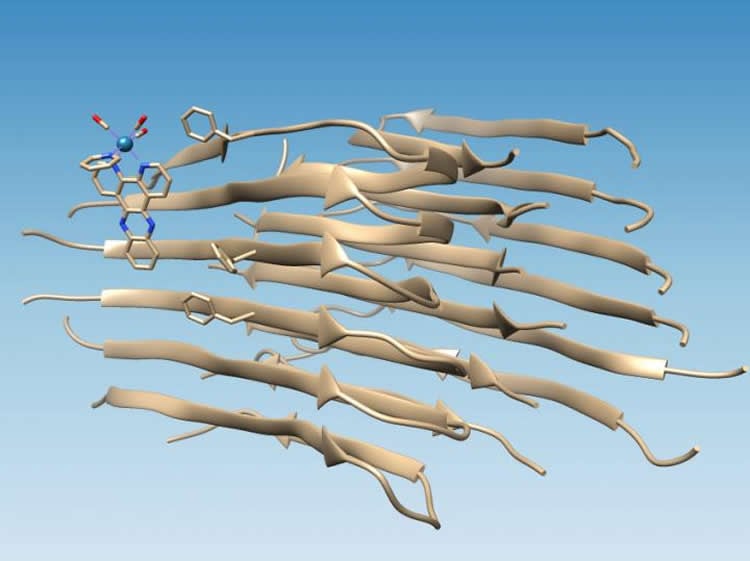
If the resulting oxidation keeps the fibrils from aggregating farther into the sticky substance found in the brains of Alzheimer’s patients, it may be the start of a useful strategy to stop aggregation before symptoms of the disease appear.
“It’s a very attractive system because it uses light, which is a cheap resource,” Martí said. “If we can modify complexes so they absorb red light, which is transparent to tissue, we might be able to perform these photochemical modifications in living animals, and maybe someday in humans.”
He said light activation allows the researchers to have “exquisite control” of oxidation.
“We imagine it might be possible someday to prevent symptoms of Alzheimer’s by targeting amyloid beta in the same way we treat cholesterol in people now to prevent cardiovascular disease,” Martí said. “That would be wonderful.”
The Welch Foundation and National Science Foundation supported the research. The Center of Computational Science at the University of Miami provided computational resources.
Source: David Ruth – Rice University
Publisher: Organized by NeuroscienceNews.com.
Image Source: NeuroscienceNews.com image is credited to Martí Group/Rice University.
Original Research: Abstract for “Photochemical Identification of Molecular Binding Sites on the Surface of Amyloid-β Fibrillar Aggregates” by Amir Aliyan, Thomas J. Paul, Bo Jiang, Christopher Pennington, Gaurav Sharma, Rajeev Prabhakar, and Angel A. Martí in Chem. Published online October 19 2017 doi:10.1016/j.chempr.2017.09.011
[cbtabs][cbtab title=”MLA”]Rice University “Binding Sites on Amyloid Beta Peptide Discovered.” NeuroscienceNews. NeuroscienceNews, 19 October 2017.
<https://neurosciencenews.com/amyloid-beta-peptide-binding-7773/>.[/cbtab][cbtab title=”APA”]Rice University (2017, October 19). Binding Sites on Amyloid Beta Peptide Discovered. NeuroscienceNews. Retrieved October 19, 2017 from https://neurosciencenews.com/amyloid-beta-peptide-binding-7773/[/cbtab][cbtab title=”Chicago”]Rice University “Binding Sites on Amyloid Beta Peptide Discovered.” https://neurosciencenews.com/amyloid-beta-peptide-binding-7773/ (accessed October 19, 2017).[/cbtab][/cbtabs]
Abstract
Photochemical Identification of Molecular Binding Sites on the Surface of Amyloid-β Fibrillar Aggregates
Highlights
•A rhenium dipyridophenazine carbonyl complex binds to Aβ fibrils
•Molecular dynamics simulations predict that binding occurs at the Phe20-Val18 cleft
•Photoirradiation of the complex causes oxidation on the Aβ fibril
•MS-MS experiments show oxidation at Met 35, consistent with Phe20-Val18 binding
The Bigger Picture
Alzheimer’s disease is a form of dementia affecting over 44 million people worldwide, and its symptoms include agitation, confusion, and memory loss. This disease is characterized by aggregates of the amyloid-β (Aβ) peptide in the brain. The transition of Aβ from the soluble to the aggregated form is linked to the onset of Alzheimer’s disease. Molecules that inhibit Aβ aggregation or quench its harmful effect are highly sought after. However, how molecules bind to Aβ is still uncertain. Aβ aggregates are disordered in nature, preventing the use of common methods for studying structure and binding. To address this, we used a rhenium complex that binds to Aβ. Upon light exposure, this complex produces oxidation on Aβ, leaving a mark at the place of binding. Spectroscopic and computational studies allowed elucidation of locations and binding modes of these molecules on Aβ. This information will guide the production of potent drugs with better binding affinities to Aβ for the treatment of Alzheimer’s disease.
Summary
The aggregation of amyloid-β (Aβ) into insoluble fibrils has been associated with the development of Alzheimer’s disease. The study of Aβ aggregation with [Re(CO)3(dppz)(Py)]+ (dppz = dipyrido[3,2-a:2′,3′-c]phenazine; Py = pyridine) has led to the observation of an irradiation-induced light-switching response accompanied by the oxidation of the Aβ fibril. Here, we used the photophysical and photochemical properties of this complex, as well as spectroscopic and computational methods, to elucidate molecular binding sites on Aβ fibrils. [Re(CO)3(dppz)(Py)]+ binds to Aβ fibrils with a dissociation constant of 4.2 μM and a binding stoichiometry 2.8:1 (Aβ/complex). Molecular dynamics (MD) simulations predicted binding of [Re(CO)3(dppz)(Py)]+ through a hydrophobic cleft on the fibril axis between Val18 and Phe20. Tandem mass spectrometry analysis indicated that oxidation occurred at Met35, footprinting the place of binding, which is close to the site predicted by the MD simulations. Finding binding sites in Aβ is of great importance for the design of Aβ-binding drugs.
“Photochemical Identification of Molecular Binding Sites on the Surface of Amyloid-β Fibrillar Aggregates” by Amir Aliyan, Thomas J. Paul, Bo Jiang, Christopher Pennington, Gaurav Sharma, Rajeev Prabhakar, and Angel A. Martí in Chem. Published online October 19 2017 doi:10.1016/j.chempr.2017.09.011