Summary: Researchers have successfully created the most detailed map of the hippocampus, an area of the brain implicated in memory. The map shows the structure, neural connections and functions in vivid detail.
Source: USC.
Curing some of the most vexing diseases first requires navigating the world’s most complex structure – the human brain. So, USC scientists have created the most detailed atlas yet of the brain’s memory bank.
Cartographers of the cranium, a USC research team has illustrated the internal circuitry of the hippocampus in detail that would make the great ancient mapmaker, Ptolemy, proud. Using fluorescent tracers and 3-D animation, the scientists show structures, nerve connections and functions in vivid detail. The study appears today in the journal Nature Neuroscience.
“Like a new atlas, we’ve constructed the most detailed diagram of the hippocampus to date,” said Michael S. Bienkowski, lead author of the study and a researcher at the USC Institute for Neuroimaging and Informatics in the Keck School of Medicine of USC. “With a better map, we can see each region and how it functions. A better map is a resource scientists can use to better understand the hippocampus and how its degeneration leads to diseases.”
The human hippocampus sits at the base of the brain and it’s shaped like a seahorse. It stores memories, helps regulate emotions and guides navigation by spatial processing. It’s the first part of the brain impaired by Alzheimer’s and hippocampus degeneration can cause epilepsy and other diseases.
In this case, scientists worked on a mouse brain because it’s organized similar to a human brain. Scientists can use the new map of the hippocampus to deliver genetically-targeted drugs to specific neurons with fewer side effects, said senior author, Hong-Wei Dong, USC professor of neurology and director of the USC Center for Integrated Connectomics (CIC).
The work is part of the Mouse Connectome Project, a USC-led effort that collects lots of data about neural connections in the brain and shares it publicly with researchers in more than 100 countries. Disconnections in the brain underlie Huntington’s disease, Parkinson’s disease, Alzheimer’s disease, autism spectrum disorders and many other illnesses.
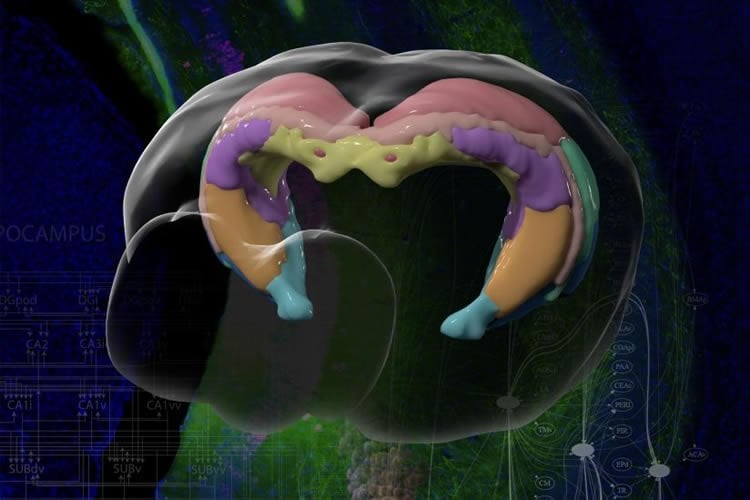
Scientists have known the basic four-part architecture of the hippocampus for a long time. What’s different now is the USC scientists can show its sub-regions and how nerve cells interact across the structure. It’s a night-and-day difference, akin to seeing transmission lines and power poles slung across a city by day compared to fully illuminated at night. This new visualization traces neural pathways and connections in remarkable detail using fluorescent dyes as tracers that reveal cells, neuron junctions and connections to the rest of the brain.
“It totally changes our understanding by combining a wiring diagram with gene expression of the mouse hippocampus. We see it doing different things, and this gives us a new way to understand how the whole thing works together. This should have a very profound and broad impact,” Bienkowski said.
Alzheimer’s disease is the sixth leading cause of death for older people and the leading cause of dementia in the United States, according to the National Institute on Aging. It accounts for 93,500 deaths nationwide annually, and the prevalence and rate of death is increasing as the population ages, according to the U.S. Centers for Disease Control and Prevention.
The Mouse Connectome Project at USC is part of a larger effort funded by the National Institutes of Health to map all the connections of the brain to understand how different structures network to regulate behavior. Dong’s research focuses on how to classify neurons based on both genetics and connectivity, information that could help other researchers develop strategies to target neurons to treat diseases in humans. The work is parallel to a Human Connectome Project, involving 100 researchers at leading research institutions with $40 million from the National Institutes of Health, that maps brain connections in humans.
The human brain contains about 100 billion neurons, each with about 10,000 connections, so mapping this network is a big-data challenge involving many scientists. USC’s contribution helps advance the larger effort by starting with a simpler task such as the mouse hippocampus research.
The study authors are Michael S. Bienkowski, Ian Bowman, Monica Y. Song, Lin Gou, Tyler Ard, K. Cotter, Muye Zhu, Nora L. Benavidez, Seita Yamashita, Jaspar Abu-Jaber, Sana Azam, Darrick Lo, Nicholas N. Foster and Houri Hintiryan of the USC Stevens Neuroimaging and Informatics Institute (INI) and Center for Integrated Connectomics (CIC) at the Keck School of Medicine; and Hong-Wei Dong of INI and CIC and the departments of physiology and neuroscience as well as the Zilkha Neurogenetic Institute at the Keck School of Medicine.
Funding: The work was supported by the National Institutes of Health/National Institute of Mental Health (R01 MH094360-01A1), NIH/NCI (U01 CA198932-01), NIH/NIMH (RF1 MH114112-01), NIH/NIMH (U01 MH114829-0), NIH/NIMH (F32 MH107071-01-A1), and a pilot fund of the Alzheimer’s Disease Research Centers (P50-AG05142).
Source: Gary Polakovic – USC
Publisher: Organized by NeuroscienceNews.com.
Image Source: NeuroscienceNews.com image is credited to Tyler Ard, USC Mark and Mary Stevens Neuroimaging and Informatics Institute.
Original Research: Abstract for “Integration of gene expression and brain-wide connectivity reveals the multiscale organization of mouse hippocampal networks” by Michael S. Bienkowski, Ian Bowman, Monica Y. Song, Lin Gou, Tyler Ard, Kaelan Cotter, Muye Zhu, Nora L. Benavidez, Seita Yamashita, Jaspar Abu-Jaber, Sana Azam, Darrick Lo, Nicholas N. Foster, Houri Hintiryan & Hong-Wei Dong in Nature Neuroscience. Published October 8 2018.
doi:10.1038/s41593-018-0241-y
[cbtabs][cbtab title=”MLA”]USC”New Map of Brain Region Linked to Alzheimer’s.” NeuroscienceNews. NeuroscienceNews, 8 October 2018.
<https://neurosciencenews.com/alzheimers-brain-map-9977/>.[/cbtab][cbtab title=”APA”]USC(2018, October 8). New Map of Brain Region Linked to Alzheimer’s. NeuroscienceNews. Retrieved October 8, 2018 from https://neurosciencenews.com/alzheimers-brain-map-9977/[/cbtab][cbtab title=”Chicago”]USC”New Map of Brain Region Linked to Alzheimer’s.” https://neurosciencenews.com/alzheimers-brain-map-9977/ (accessed October 8, 2018).[/cbtab][/cbtabs]
Abstract
Integration of gene expression and brain-wide connectivity reveals the multiscale organization of mouse hippocampal networks
Understanding the organization of the hippocampus is fundamental to understanding brain function related to learning, memory, emotions, and diseases such as Alzheimer’s disease. Physiological studies in humans and rodents have suggested that there is both structural and functional heterogeneity along the longitudinal axis of the hippocampus. However, the recent discovery of discrete gene expression domains in the mouse hippocampus has provided the opportunity to re-evaluate hippocampal connectivity. To integrate mouse hippocampal gene expression and connectivity, we mapped the distribution of distinct gene expression patterns in mouse hippocampus and subiculum to create the Hippocampus Gene Expression Atlas (HGEA). Notably, previously unknown subiculum gene expression patterns revealed a hidden laminar organization. Guided by the HGEA, we constructed the most detailed hippocampal connectome available using Mouse Connectome Project (http://www.mouseconnectome.org) tract tracing data. Our results define the hippocampus’ multiscale network organization and elucidate each subnetwork’s unique brain-wide connectivity patterns.






