Age may seem like a straightforward measurement: the number of years, months, days since you were born. But for cells in different parts of your body, age can mean very different things. Scientists at the European Molecular Biology Laboratory (EMBL) in Heidelberg, Germany, and the Salk Institute and the University of California at Berkeley, both in the USA, have now measured and compared just how ageing affects rats’ liver and brain cells. In a study published online today in Cell Systems, they were able to tease out general ageing processes from those that are specific to each of these organs.
“We found that in brain, age-related changes very often have to do with the loss of molecules that help signals to spread among neurons. This could explain why old rats have a reduced ability to form new connections between neurons, as well as other traits observed in the aging brain,” says Martin Beck, who led the work at EMBL. “And it is very similar to what has been found in previous studies that have looked at gene expression in humans.”
The scientists compared brain and liver cells of rats in the prime of life – 6 months old, roughly equivalent to 18 year-old humans – to those of old rats – 24 months old. Rather than focus solely on gene expression – which genes are turned on or off – as most previous studies had done, the team employed a variety of techniques to assess several steps in the cells’ protein production assembly line. They measured which parts of the genome had been transcribed into RNA, what proteins the cells had produced, what rates proteins were being produced at, and what chemical markers were added to proteins in ‘post-production’.
“Integrating data from different levels was crucial,” says Beck. “When you see how the data sets cluster, it reveals what’s actually going on. Often it was only then that we could see that a whole network of reactions is affected.”
Ageing impacted some networks equally in liver and brain – notably immune response and inflammation, and stress responses – implying that these are probably general effects of ageing, felt throughout the body. But other effects were very specific. The livers of old rats showed changes in metabolic processes – i.e. how cells process molecules – mostly at the level of regulating how the genome is ‘read’. In the brain, by contrast, the tell-tale signs of ageing were mostly at the level of protein production, and mainly affected signalling processes that enable neurons to communicate with each other.
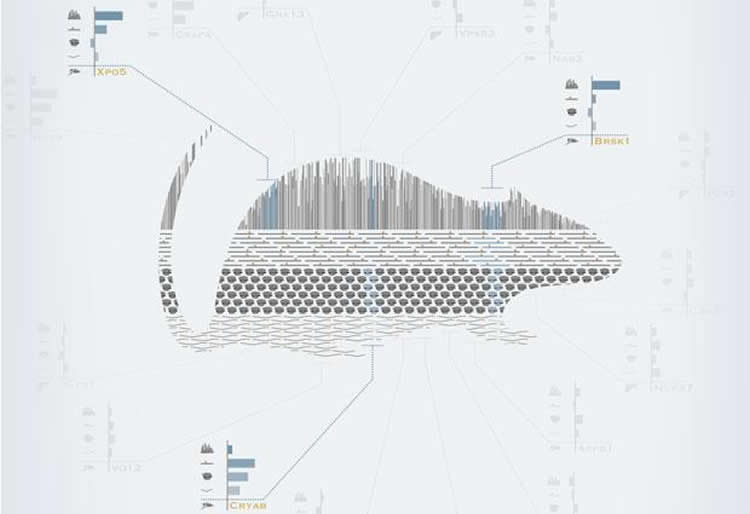

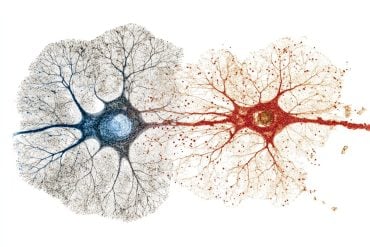
“We chose to compare brain and liver because these two organs have very different capacities for self-renewal,” says Martin Hetzer, who led the work at the Salk Institute. “The liver has a very high regeneration rate, so even in an old animal – or person – most of the molecules in liver cells will actually be relatively young. In the brain, it’s the opposite: in our previous work we have shown that many sets of molecules in the brain are effectively there for life.”
“This is just the beginning,” says Nicholas Ingolia, who led the work at Berkeley. “Our study provides a rather large resource of proteins, RNAs and post-translational modifications that are affected by age, which we hope others can use to propose new hypotheses which can be tested experimentally.”
Funding: This research is funded by Alexander von Humboldt foundation, Marie Curie Actions, EMBL Interdisciplinary Postdoc Programme, Marie Curie Actions COFUND, Searle Scholars Program, and Paul F. Glenn Center for Aging Research.
Source: Sonia Furtado Neves – EMBL
Image Credit: The image is credited to Brandon Toyama/EMBL
Original Research: Full open access research for “Integrated Transcriptome and Proteome Analyses Reveal Organ-Specific Proteome Deterioration in Old Rats” by Alessandro Ori, Brandon H. Toyama, Michael S. Harris, Thomas Bock, Murat Iskar, Peer Bork, Nicholas T. Ingolia, Martin W. Hetzer, and Martin Beck in Cell Systems. Published online September 17 2015 doi:10.1016/j.cels.2015.08.012
Abstract
Integrated Transcriptome and Proteome Analyses Reveal Organ-Specific Proteome Deterioration in Old Rats
Highlights
•An integrated approach identifies molecular alterations between young and old rats
•Changes in translation output explain the majority of the altered protein abundances
•Key protein complexes are altered in abundance and composition
•We provide a rich data resource to stimulate further studies of aging
Summary
Aging is associated with the decline of protein, cell, and organ function. Here, we use an integrated approach to characterize gene expression, bulk translation, and cell biology in the brains and livers of young and old rats. We identify 468 differences in protein abundance between young and old animals. The majority are a consequence of altered translation output, that is, the combined effect of changes in transcript abundance and translation efficiency. In addition, we identify 130 proteins whose overall abundance remains unchanged but whose sub-cellular localization, phosphorylation state, or splice-form varies. While some protein-level differences appear to be a generic property of the rats’ chronological age, the majority are specific to one organ. These may be a consequence of the organ’s physiology or the chronological age of the cells within the tissue. Taken together, our study provides an initial view of the proteome at the molecular, sub-cellular, and organ level in young and old rats.
“Evidence for Multiple Rhythmic Skills” by Adam Tierney and Nina Kraus in PLOS ONE. Published online September 16 2015 doi:10.1371/journal.pone.0136645.g001