Summary: Researchers join their own experiments as test subjects to evaluate different brain networks. Using neuroimaging to scan their brains, researchers created individual detailed maps of neural connections.
Source: WUSTL.
Unique features of individual brains captured.
A quest to analyze the unique features of individual human brains evolved into the so-called Midnight Scan Club, a group of scientists who had big ideas but almost no funding and little time to research the trillions of neural connections that activate the body’s most powerful organ.
The research group started in 2013 by two neuroscientists at Washington University School of Medicine in St. Louis who aimed to collect a massive amount of data on individual brains. The study’s subjects were the scientists themselves and eight others, all junior faculty or graduate students.
Most efforts to analyze connections involve scanning many brains and averaging the data across groups of people. For this study, the researchers used brain-imaging techniques to evaluate brain networks that control speech and motor function, among other activities. The researchers examined individuals while resting and performing cognitive tasks such as reading.
Their aim was to better understand the inner-workings of individual people’s brains by compiling hours and hours of data collected on 10 adults — a group of study subjects that included two of the researchers themselves. Their work led to 10 high-fidelity, individual-specific connectomes — detailed maps of neural brain connections that reveal spatial and organizational variability in brain networks, and that may one day be helpful in determining personalized treatments for brain-related disorders.
The research is published online July 27 in the journal Neuron and will be featured on the cover of the Aug. 16 print edition of journal. The dataset is available as a resource for neuroscientists so they can plumb the information for further discoveries about brain circuitry.
“Using this approach, we might one day be able to obtain brain-function maps of patients and individualize neuropsychiatric and medical treatments based on specific features of their brain networks,” said Nico Dosenbach, MD, PhD, an assistant professor of pediatric and developmental neurology at the School of Medicine. “We’d like to be able to provide personalized medicine using neuroimaging to treat people with seizures, migraines and depression.
“This is also exciting for basic neuroscience because these individual maps look quite a bit different from the currently available standard group maps,” Dosenbach said. “We learned that spatial averaging destroys all the fine features of our brain organization, kind of like what happens when you average photos of several peoples’ faces. Everything gets blurred and blotchy.”
Over the past 25 years, neurologists have averaged MRI data across groups of people as a way to approximate human brain organization.
“We wanted to take a different approach,” explained Dosenbach, who also treats patients at St. Louis Children’s Hospital. “Averaging brain data across many individuals has allowed scientists to learn a lot about brain function. But compiling data on an individual level invites opportunities for precision. The goal is to find individual variations that may play a role in neurological diseases, from migraines to Alzheimer’s to brain injuries.”
Dosenbach and his colleagues collected MRI images of 10 individuals and analyzed how nerve cells function in individual brains. They used two types of magnetic resonance imaging: functional MRI, which pinpoints the parts of the brain most active during specific cognitive activities; and resting-state functional connectivity, to explore neural connections while the brain is not actively engaged in a task.
Getting such images, however, presented challenges — among them, finding committed volunteers to undergo hours of loud, unpleasant brain imaging, and finding a way around the high demand for and cost of MRI machines for the research.
“The major downside of getting massive amounts of imaging data per person is its great cost,” Dosenbach said. “And as a new faculty member, I was extremely short on funding. In addition, the scanner was mostly booked throughout the day, every day.”
Undeterred, Dosenbach and his research partner, Steve Nelson, PhD, then a research scientist in psychology at Washington University, brainstormed for weeks before Nelson discovered that the fee for one of the university’s research scanners, which at the time was about $600 per hour, was reduced by 90 percent after midnight.
“Our motto became ‘Carpe noctem,’ or ‘Seize the night,’ ” Dosenbach said.
As founding members of the Midnight Scan Club, Dosenbach and Nelson — now a neuroimaging director at the U.S. Department of Veterans Affairs in Waco, Texas, and a research assistant professor at Baylor University in Waco and the University of Texas at Dallas — scanned their brains first, subjecting themselves to 12 nightly, two-hour sessions.
The two then recruited eight volunteers — male and female graduate students and junior-level faculty in their mid-20s to mid-30s.
“The study wasn’t easy,” Dosenbach said. “We were already stretched thin professionally, and the MRI experience was unpleasant and it was hard to stay awake. And we had personal lives, too. At the time, I had a toddler, and my wife was seven months pregnant. It was a big sacrifice for her to have me gone for much of the day and the night, and then for me to be napping when I was home.”
However, the strategy proved budget-friendly, costing a total of $12,000 instead of about $125,000 had they performed the scans during the day.
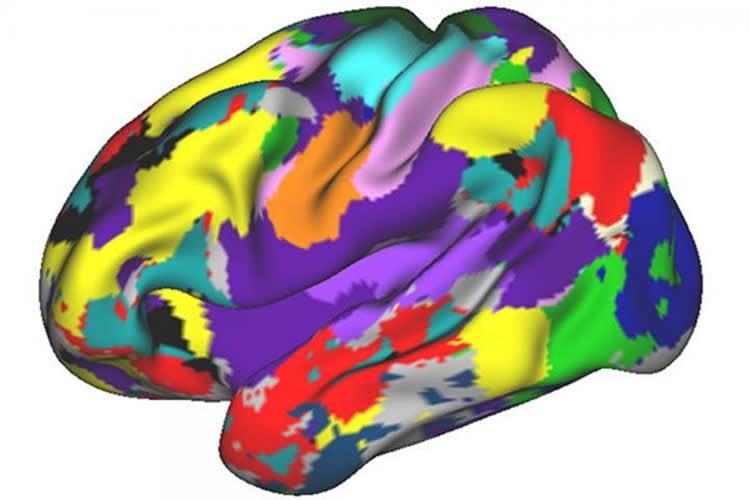
Their analysis — led by first author Evan Moss Gordon, PhD, an investigator at the U.S. Department of Veterans Affairs in Waco, and Timothy O. Laumann, MD, PhD, a resident in psychiatry at Barnes-Jewish Hospital — revealed several variations in brain organization among the group. For example, in most individuals the entire brain network forms a giant loop; however, in two of the study participants, including Dosenbach, the brain as a whole showed more linear connections.
“What does it mean that my brain is different?” he asked. “I don’t know, however, it’s generally believed that a circular network increases efficiency of information processing.”
He compared the scenario to two drivers who choose different routes to the same destination. One hops on the freeway and arrives quickly; the other meanders the side streets. Both arrive at the same place despite different methods and time frames.
“In future work with larger sample sizes, we hope to clarify which variations in neural connections affect behavior and which do not,” Dosenbach said.
The dataset also may help neurologists understand and treat brain disorders such as seizures, Dosenbach said. “If we collect more and higher-quality data per subject, we can make precise maps of each person’s functional brain architecture,” he said. “To me, that’s step one toward using those maps to guide us in the care of neurological and psychiatric patients. Our small sample is not diverse, yet the data already has suggested several variations. This may be important for understanding why people respond uniquely to different drug treatments.”
The project was inspired, in part, by the Human Connectome Project, a global effort focused on mapping human brain circuitry through neuroimaging and data-sharing. Washington University has co-led the project since 2010, when the National Institutes of Health (NIH) named the university a co-leader in overseeing $30 million to a research consortium to fulfill the grant.
Dosenbach also said he was inspired by Russell Poldrack, PhD, a noted neuroscientist and psychologist at Stanford University. “Russ was the first to scan himself twice a week for almost 18 months,” Dosenbach said. “We admired his approach and we wanted to expand on it.
“We’ve already had other research groups express interest in mimicking our data-collecting methods, and other research groups want to collaborate on analyzing the data,” he said. “For me, this is a paradigm shift in the field of neuroimaging. I am excited to see where this will lead us in our quest to uncover the mysteries of the human brain.”
Funding: The study was supported by National Institutes of Health, Jacobs Foundation, Child Neurology Foundation, McDonnell Center for Systems Neuroscience, Mallinckrodt Institute of Radiology, Hope Center for Neurological Disorders, Dart Neuroscience LLC.
The authors state that they have no competing financial interests.
Source: Diane Duke Williams – WUSTL
Image Source: NeuroscienceNews.com image is credited to Washington University.
Original Research: Abstract for “Precision Functional Mapping of Individual Human Brains” by Evan M. Gordon, Timothy O. Laumann, Adrian W. Gilmore, Dillan J. Newbold, Deanna J. Greene, Jeffrey J. Berg, Mario Ortega, Catherine Hoyt-Drazen, Caterina Gratton, Haoxin Sun, Jacqueline M. Hampton, Rebecca S. Coalson, Annie L. Nguyen, Kathleen B. McDermott, Joshua S. Shimony, Abraham Z. Snyder, Bradley L. Schlaggar, Steven E. Petersen, Steven M. Nelson,and Nico U.F. Dosenbach in Neuron. Published online July 27 2017 doi:10.1016/j.neuron.2017.07.011
[cbtabs][cbtab title=”MLA”]WUSTL “Scientists Become Research Subjects in After-Hours Brain-Scanning Project.” NeuroscienceNews. NeuroscienceNews, 27 July 2017.
<research-subjects-neuroscience-7190/>.[/cbtab][cbtab title=”APA”]WUSTL (2017, July 27). Scientists Become Research Subjects in After-Hours Brain-Scanning Project. NeuroscienceNew. Retrieved July 27, 2017 from research-subjects-neuroscience-7190/[/cbtab][cbtab title=”Chicago”]WUSTL “Scientists Become Research Subjects in After-Hours Brain-Scanning Project.” research-subjects-neuroscience-7190/ (accessed July 27, 2017).[/cbtab][/cbtabs]
Abstract
Precision Functional Mapping of Individual Human Brains
Highlights
•Individual brain organization is qualitatively different from group-average estimates
•Individualized measures of brain function become reliable with large amounts of data
•Individuals exhibit distinct brain network topography and topology
•We release highly sampled, multi-modal fMRI data on ten subjects as a NeuroResource
Summary
Human functional MRI (fMRI) research primarily focuses on analyzing data averaged across groups, which limits the detail, specificity, and clinical utility of fMRI resting-state functional connectivity (RSFC) and task-activation maps. To push our understanding of functional brain organization to the level of individual humans, we assembled a novel MRI dataset containing 5 hr of RSFC data, 6 hr of task fMRI, multiple structural MRIs, and neuropsychological tests from each of ten adults. Using these data, we generated ten high-fidelity, individual-specific functional connectomes. This individual-connectome approach revealed several new types of spatial and organizational variability in brain networks, including unique network features and topologies that corresponded with structural and task-derived brain features. We are releasing this highly sampled, individual-focused dataset as a resource for neuroscientists, and we propose precision individual connectomics as a model for future work examining the organization of healthy and diseased individual human brains.
“Precision Functional Mapping of Individual Human Brains” by Evan M. Gordon, Timothy O. Laumann, Adrian W. Gilmore, Dillan J. Newbold, Deanna J. Greene, Jeffrey J. Berg, Mario Ortega, Catherine Hoyt-Drazen, Caterina Gratton, Haoxin Sun, Jacqueline M. Hampton, Rebecca S. Coalson, Annie L. Nguyen, Kathleen B. McDermott, Joshua S. Shimony, Abraham Z. Snyder, Bradley L. Schlaggar, Steven E. Petersen, Steven M. Nelson,and Nico U.F. Dosenbach in Neuron. Published online July 27 2017 doi:10.1016/j.neuron.2017.07.011






