Summary: Researchers have built a decoder which is able to translate neural signals into mood variations.
Source: USC.
By developing a novel decoding technology, a team of engineers and physicians at the University of Southern California (USC) and UC San Francisco have discovered how mood variations can be decoded from neural signals in the human brain–a process that has not been demonstrated to date.
Their study, published in Nature Biotechnology, is a significant step towards creating new closed-loop therapies that use brain stimulation to treat debilitating mood and anxiety disorders in millions of patients who are not responsive to current treatments.
Assistant Professor and Viterbi Early Career Chair Maryam Shanechi of the Ming Hsieh Department of Electrical Engineering and the Neuroscience Graduate Program at USC led the development of the decoding technology, and Professor of Neurological Surgery Edward Chang at UCSF led the human implantation and data collection effort. The researchers were supporting the Defense Advanced Research Projects Agency’s SUBNETS program to develop new biomedical technologies for treating intractable neurological diseases.
The team recruited seven human volunteers among a group of epilepsy patients who already had intracranial electrodes inserted in their brain for standard clinical monitoring to locate their seizures. Large-scale brain signals were recorded from these electrodes in the volunteers across multiple days at UCSF, while they also intermittently reported their moods using a questionnaire. Shanechi and her students, Omid Sani and Yuxiao Yang, used that data to develop a novel decoding technology that could predict mood variations over time from the brain signals in each human subject, a goal unachievable to date.
“Mood is represented across multiple sites in the brain rather than localized regions, thus decoding mood presents a unique computational challenge,” Shanechi said. “This challenge is made more difficult by the fact that we don’t have a full understanding of how these regions coordinate their activity to encode mood and that mood is inherently difficult to assess. To solve this challenge, we needed to develop new decoding methodologies that incorporate neural signals from distributed brain sites while dealing with infrequent opportunities to measure moods.”
To build the decoder, Shanechi and the team of researchers analyzed brain signals that were recorded from intracranial electrodes in the seven human volunteers. Raw brain signals were continuously recorded across distributed brain regions while the patients self-reported their moods through a tablet-based questionnaire.
In each of the 24 questions, the patient was asked to “rate how you feel now” by tapping one of 7 buttons on a continuum between a pair of negative and positive mood state descriptors (e.g., “depressed” and “happy”). A higher score corresponded to a more positive mood state.
Using their methodology, the researchers were able to uncover the patterns of brain signals that matched the self-reported moods. They then used this knowledge to build a decoder that would independently recognize the patterns of signals corresponding to a certain mood. Once the decoder was built, it measured the brain signals alone to predict mood variations in each patient over multiple days.
A Potential Solution for Untreatable Neuropsychiatric Conditions?
The USC/UCSF team believe their findings could support the development of new closed-loop brain stimulation therapies for mood and anxiety disorders.
Data from the 2016 National Survey on Drug Use and Health revealed that 16.2 million adults in the United States (approximately 6.7 % of all U.S. adults) have suffered at least one major depressive episode. Treatments such as selective serotonin reuptake inhibitors (SSRIs) can be effective in some but not all patients.
According to the National Institutes of Health-funded STAR*D trial–the longest study to evaluate depression treatments–almost 33% of major depression patients do not respond to treatment (more than 5.3M people in the U.S. alone). Also, in June 2018, Centers for Disease Control and Prevention reported that suicide is on the rise in the U.S.
For the millions of treatment-resistant patients, alternative therapies may be effective. For example, human imaging studies using positron emission tomography (PET) and functional magnetic resonance imaging (fMRI) have suggested that several brain regions mediate depression, and thus brain stimulation therapies in which a mood-relevant region is electrically stimulated may be applied to alleviate depressive symptoms. While open-loop brain stimulation treatments hold some promise, a more precise, effective therapy could necessitate a closed-loop approach in which an objective tracking of mood over time guides how stimulation is delivered.
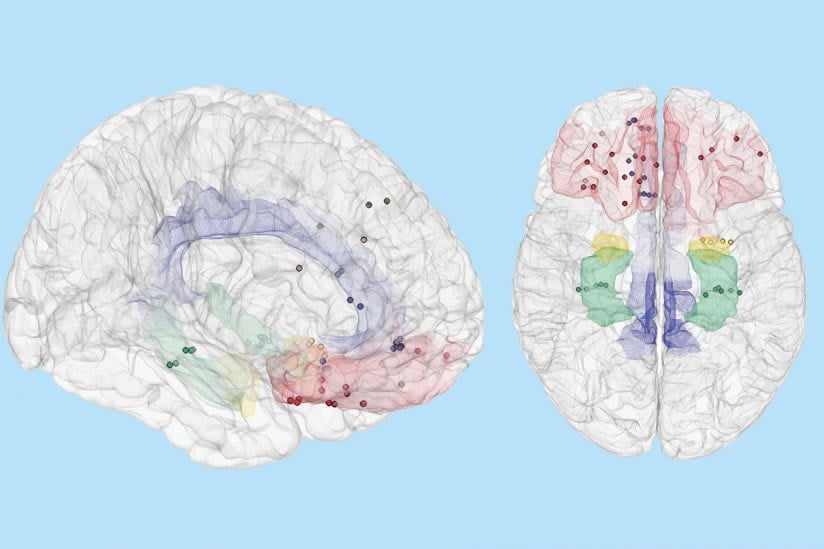
According to Shanechi, for clinical practitioners, a powerful decoding tool would provide the means to clearly delineate, in real time, the network of brain regions that support emotional behavior.
“Our goal is to create a technology that helps clinicians obtain a more accurate map of what is happening in a depressed brain at a particular moment in time and a way to understand what the brain signal is telling us about mood. This will allow us to obtain a more objective assessment of mood over time to guide the course of treatment for a given patient,” Shanechi said. “For example, if we know the mood at a given time, we can use it to decide whether or how electrical stimulation should be delivered to the brain at that moment to regulate unhealthy, debilitating extremes of emotion. This technology opens the possibility of new personalized therapies for neuropsychiatric disorders such as depression and anxiety for millions who are not responsive to traditional treatments.”
The new decoding technology, Shanechi explained, could also be extended to develop closed-loop systems for other neuropsychiatric conditions such as chronic pain, addiction, or post-traumatic stress disorder whose neural correlates are again not anatomically localized, but rather span a distributed network of brain regions, and whose behavioral assessment is difficult and thus not frequently available.
Funding: This research was partially funded by the Defense Advanced Research Projects Agency (DARPA) under Cooperative Agreement Number W911NF-14-2-0043, issued by the Army Research Office contracting office in support of DARPA’S SUBNETS program. The views, opinions, and/or findings expressed are those of the author(s) and should not be interpreted as representing the official views or policies of the Department of Defense or the U.S. Government.
Source: Amy Blumenthal – USC
Publisher: Organized by NeuroscienceNews.com.
Image Source: NeuroscienceNews.com image is credited to Sani et. al., Nature Biotechnology.
Original Research: Abstract for “Mood variations decoded from multi-site intracranial human brain activity” by Omid G Sani, Yuxiao Yang, Morgan B Lee, Heather E Dawes, Edward F Chang & Maryam M Shanechi in Nature Biotechnology. Published September 10 2018.
doi:10.1038/nbt.4200
[cbtabs][cbtab title=”MLA”]USC”Decoding Mood from Human Brain Signals.” NeuroscienceNews. NeuroscienceNews, 10 September 2018.
<https://neurosciencenews.com/mood-brain-signals-9833/>.[/cbtab][cbtab title=”APA”]USC(2018, September 10). Decoding Mood from Human Brain Signals. NeuroscienceNews. Retrieved September 10, 2018 from https://neurosciencenews.com/mood-brain-signals-9833/[/cbtab][cbtab title=”Chicago”]USC”Decoding Mood from Human Brain Signals.” https://neurosciencenews.com/mood-brain-signals-9833/ (accessed September 10, 2018).[/cbtab][/cbtabs]
Abstract
Mood variations decoded from multi-site intracranial human brain activity
The ability to decode mood state over time from neural activity could enable closed-loop systems to treat neuropsychiatric disorders. However, this decoding has not been demonstrated, partly owing to the difficulty of modeling distributed mood-relevant neural dynamics while dealing with the sparsity of mood state measurements. Here we develop a modeling framework to decode mood state variations from multi-site intracranial recordings in seven human subjects with epilepsy who self-reported their mood state intermittently over multiple days. We built dynamic neural encoding models of mood state and corresponding decoders for each individual and demonstrated that mood state variations over time can be decoded from neural activity. Across subjects, the decoders largely recruited neural signals from limbic regions, whose spectro-spatial features were tuned to mood variations. The dynamic models also provided an analytical tool to compute the timescales of the decoded mood state. These results provide an initial line of evidence indicating the feasibility of mood state decoding.