Researchers at Harvard Medical School and Dana-Farber Cancer Institute report that humans have a surprisingly high number of genes in which one of two copies has been randomly silenced. They suggest that the phenomenon is an important source of genetic diversity among people and also may help explain the genetic variation among cancer cells in a tumor.
A team led by Alex Gimelbrant, HMS assistant professor of genetics at Dana-Farber, reporting in Nature Genetics, found that genes with monoallelic expression, or MAE, contribute disproportionately to human genetic diversity.
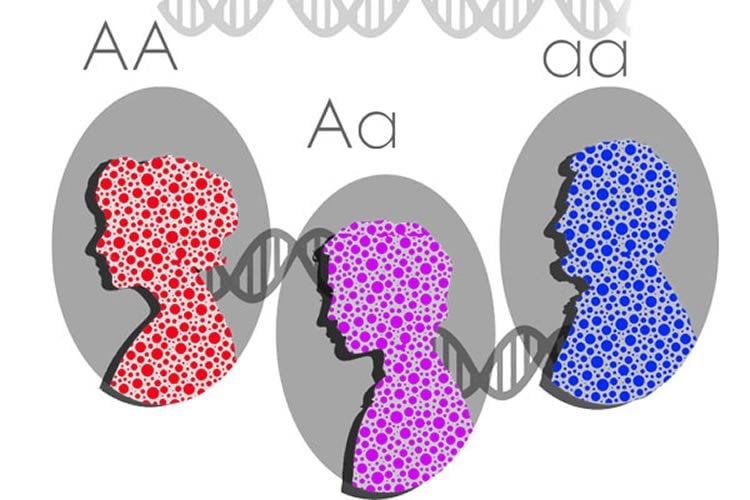
The human genome includes between 20,000 and 25,000 genes that encode the recipes for making proteins. Each gene in a cell exists in two copies, or alleles, one inherited from the father and one from the mother. The copies may be identical or different, but in most cases both alleles are expressed—actively issuing instructions for making a protein. When only one of a pair of alleles is active, that gene is said to have monoallelic expression.
A few examples of such genes have been known for several decades, but Gimelbrant and other scientists have more recently developed powerful tools for searching for them in the genome. Studies have estimated that MAE can occur in an estimated 10 to 25 percent of genes in many types of cells. For example, such genes are found in cells of the immune system and the olfactory system, where genetic diversity helps otherwise identical cells respond to many different combinations of stimuli.
In such genes where only one allele is expressed, it’s not because the other copy has been damaged or lost. Instead, said Gimelbrant, in a gene with monoallelic expression, there’s an equal chance for either of the two copies to be expressed during early development—but only one of the copies will be expressed. This random decision doesn’t change the DNA in that cell, but the decision to express only one allele is “inherited” by the offspring of that cell, through what is called an epigenetic process. In other words, the identical genes in the daughter cells will also be monoallelic.
In the new research, the investigators studied 4,227 genes with monoallelic expression to assess the extent of genetic variation of these genes and their contribution to human diversity. They said they were surprised to discover that these genes have unusually high genetic variation. As a result, the MAE genes “substantially increase the potential for cell-to-cell variability within an individual,” the authors wrote. “It looks like this is a very widespread mechanism that is responsible for genetic variation.” Not only widespread, but ancient: They traced this mechanism back to the Neanderthals and said it exists in chimpanzees.
From an evolutionary standpoint, individuals whose cells have genetic variability may more successfully adapt to their environment. A well-known example is that of genes causing sickle cell anemia. Individuals who carry identical genes for the disease have a shorter life expectancy than those whose cells have two normal genes. But in parts of Africa where malaria is epidemic, people who are heterozygous—one sickle cell allele and one normal allele—are more resistant to developing malaria.
Cancer tumors are made up of cells that grow and evolve much as populations do, and they are often found to be genetically heterogeneous. That is, the tumors include cells of several distinct genetic types, some of which may be vulnerable to a cancer drug while others are not. As a result, physicians need to try different drugs or combinations to prevent the tumor from escaping treatment.
“We think this variation may be a result of monoallelic expression of cells in the tumor,” said Gimelbrant.
MAE may also account for a situation in which two siblings, for example, may carry the same gene that raises the risk of cancer or some other disease, and yet one of them develops the condition while the other remains healthy.
Funding: Research support was provided by National Institutes of Health award GM114864.
Source: Richard Saltus – Harvard
Image Source: The image is credited to Victoria Tentler Krylov and Virginia Savova
Original Research: Abstract for “Genes with monoallelic expression contribute disproportionately to genetic diversity in humans” by Virginia Savova, Sung Chun, Mashaal Sohail, Ruth B McCole, Robert Witwicki, Lisa Gai, Tobias L Lenz, C-ting Wu, Shamil R Sunyaev and Alexander A Gimelbrant in Nature Genetics. Published online January 25 2016 doi:10.1038/ng.3493
Abstract
Genes with monoallelic expression contribute disproportionately to genetic diversity in humans
An unexpectedly large number of human autosomal genes are subject to monoallelic expression (MAE). Our analysis of 4,227 such genes uncovers surprisingly high genetic variation across human populations. This increased diversity is unlikely to reflect relaxed purifying selection. Remarkably, MAE genes exhibit an elevated recombination rate and an increased density of hypermutable sequence contexts. However, these factors do not fully account for the increased diversity. We find that the elevated nucleotide diversity of MAE genes is also associated with greater allelic age: variants in these genes tend to be older and are enriched in polymorphisms shared by Neanderthals and chimpanzees. Both synonymous and nonsynonymous alleles of MAE genes have elevated average population frequencies. We also observed strong enrichment of the MAE signature among genes reported to evolve under balancing selection. We propose that an important biological function of widespread MAE might be the generation of cell-to-cell heterogeneity; the increased genetic variation contributes to this heterogeneity.
“Genes with monoallelic expression contribute disproportionately to genetic diversity in humans” by Virginia Savova, Sung Chun, Mashaal Sohail, Ruth B McCole, Robert Witwicki, Lisa Gai, Tobias L Lenz, C-ting Wu, Shamil R Sunyaev and Alexander A Gimelbrant in Nature Genetics. Published online January 25 2016 doi:10.1038/ng.3493