Summary: Using a new advanced imaging technology, researchers capture new patterns of molecular organization as connections between neurons strengthen during learning tasks.
Source: Thomas Jefferson University.
When we learn the connections between neurons strengthen. Addiction or other neurological diseases are linked to abnormally strong connections. But what does learning look like on the cellular and molecular level? How do our cells change when we learn? Using super-resolution live-cell microscopy, researchers at Thomas Jefferson University zoomed into the connections between neurons that strengthen to discover structural changes that had never been seen before.
The research was published April 23rd in Nature Neuroscience.
“Our observations give the field a new way of thinking about how normal learning and the maladaptive learning we see in disorders, such as addiction or autism, might occur,” said Matthew Dalva Professor of Neuroscience at The Vickie and Jack Farber Institute for Neuroscience and Director of the Synaptic Biology Center at Jefferson (Philadelphia University + Thomas Jefferson University).
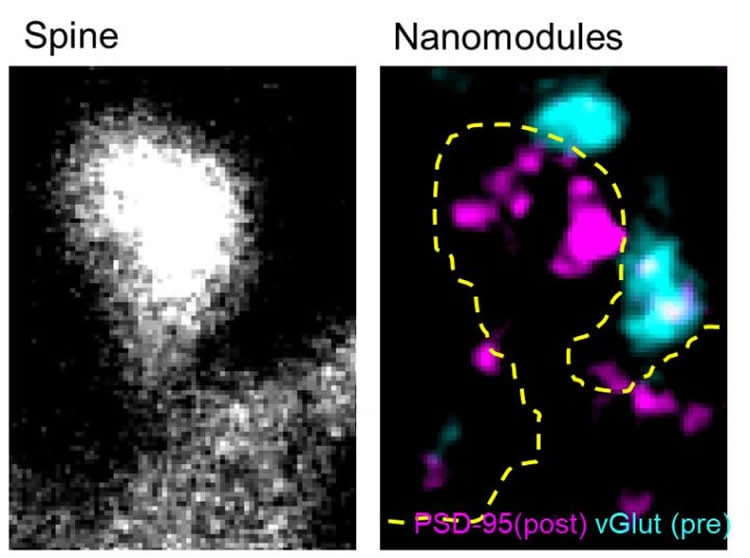
Rather than simply seeing bigger connections during learning, which has been observed before, Dr. Dalva and his colleagues found that the molecules involved in sending and receiving the signals between neurons appeared to be organized in clumps or “nanomodules” that both dance and multiply when stimulated by learning-like signals.
The researchers made their observations using living neurons in real-time, zooming into synapses, the sites of neuronal connection where information is passed from one cell to another to enable learning and other behavior. Dr. Dalva’s colleagues visualized the key molecules involved in the neurotransmission from neuron to neuron with two colors, green on sending side (the pre-synaptic side) and red on the receiving side (postsynaptic side).
The team made a number of surprising observations about the synaptic nanomodules. They saw that the key molecules on the presynaptic side clumped together and tracked, as if linked, to the key molecules clumped on the postsynaptic side. These pre/post molecular clumps or nanomodules appear to have a uniform size. They also multiplied when the neurons were stimulated in a way that mimicked changes in the size of the spines which protrude from neurons to nearly touch at the synapse. And as the number of nanomodules increased, so did the size of the spines. “The key finding is that changes in synaptic strength might be more digital than analog – with same sized units added to change synaptic strength,” said Dr. Dalva.
Another surprise was how the nanomodules behaved when stimulated. “When we activated the neuron with signals that would strengthen the synaptic connection, a non-moving nanomodule would begin to jiggle and move around the synaptic spine, with the pre- and post-synaptic components always in lock step,” said first author Dr. Martin Hruska, an Instructor in Dr. Dalva’s lab.
“Although it’s yet unclear how these nanomodules might behave in disease states, our observations offer a new way to explore those questions,” said Dr. Dalva.
Like the best scientific research, the research suggests many more questions for the field to investigate: How is the neuron able to make the clumps or nanodomains of the same size? Why are they the same size? How do they increase in number – do they split in two, or are new ones made? Why do they move around when the synapse is stimulated? Finally, how do nanomodules behave in disorders such as addiction or autism?
Funding: This research was supported by grants from NIDA (DA022727), NIMH (MH100093) and the Vickie and Jack Farber Foundation. The funders had no role in the study design, conduct of the experiments, and interpretation of the data.
Source: Edyta Zielinska – Thomas Jefferson University
Publisher: Organized by NeuroscienceNews.com.
Image Source: NeuroscienceNews.com image is credited to Matthew Dalva Lab, Jefferson (Philadelphia University + Thomas Jefferson University).
Original Research: Abstract for “Synaptic nanomodules underlie the organization and plasticity of spine synapses” by Martin Hruska, Nathan Henderson, Sylvain J. Le Marchand, Haani Jafri & Matthew B. Dalva in Nature Neuroscience. Published April 23 2018.
doi:10.1038/s41593-018-0138-9
[cbtabs][cbtab title=”MLA”]Thomas Jefferson University “What Learning Looks Like In the Brain.” NeuroscienceNews. NeuroscienceNews, 23 April 2018.
<https://neurosciencenews.com/learning-brain-8862/>.[/cbtab][cbtab title=”APA”]Thomas Jefferson University (2018, April 23). What Learning Looks Like In the Brain. NeuroscienceNews. Retrieved April 23, 2018 from https://neurosciencenews.com/learning-brain-8862/[/cbtab][cbtab title=”Chicago”]Thomas Jefferson University “What Learning Looks Like In the Brain.” https://neurosciencenews.com/learning-brain-8862/ (accessed April 23, 2018).[/cbtab][/cbtabs]
Abstract
Synaptic nanomodules underlie the organization and plasticity of spine synapses
Experience results in long-lasting changes in dendritic spine size, yet how the molecular architecture of the synapse responds to plasticity remains poorly understood. Here a combined approach of multicolor stimulated emission depletion microscopy (STED) and confocal imaging in rat and mouse demonstrates that structural plasticity is linked to the addition of unitary synaptic nanomodules to spines. Spine synapses in vivo and in vitro contain discrete and aligned subdiffraction modules of pre- and postsynaptic proteins whose number scales linearly with spine size. Live-cell time-lapse super-resolution imaging reveals that NMDA receptor–dependent increases in spine size are accompanied both by enhanced mobility of pre- and postsynaptic modules that remain aligned with each other and by a coordinated increase in the number of nanomodules. These findings suggest a simplified model for experience-dependent structural plasticity relying on an unexpectedly modular nanomolecular architecture of synaptic proteins.