Summary: Histone H4 mutations in the spool cause intellectual disabilities and developmental delays, a new study reports.
Source: University of Otago
University of Otago-led research has identified a new rare genetic mutation which will finally give families around the world answers as to what is affecting their loved one.
The mutation was first discovered in a New Zealand patient and has now been identified in 29 people in 10 countries.
Lead researcher Dr. Louise Bicknell says the finding will not only supply answers and reassurance to patients and their families, it gives greater insight into the factors required for brain function.
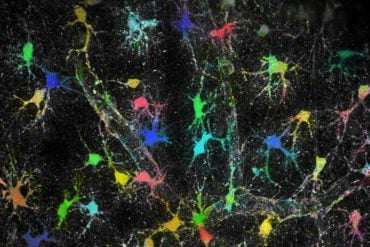
Patients with the disorder have a wide range of intellectual disability—in some it is mild, in others it much more severe, with a poorer quality of life. Some have other neurological features such as seizures, poor muscle coordination, or autism or ADHD.
“The family would have been told that their child’s condition is highly likely to have a genetic cause, but now through our research, they will have a formal diagnosis—which can help initiate more support for the family. This also means that they can get information about the chances of having another child that is affected—which is very low—which helps with family planning.”
In order for DNA to fit into a cell, it must be wound up like thread on a spool, she says.
“Until recently, the spool was often ignored and seen as just packaging material for DNA. Our genetic studies have discovered that altering this spool can disrupt brain development and functioning.
“We have found that mutations in one of the spool’s components (histone H4) causes intellectual disability and delayed development in a group of individuals from 10 different countries,” she says.
The finding started after studies on a New Zealand child.
“We were able to sequence the genes in her DNA, and identified this possible mutation, which hadn’t been seen before in other research. We then teamed up with Associate Professor Gijs van Haaften, in the Netherlands, who was also interested in this disorder—and through international networks, genetic matching websites and word of mouth, we have gathered together a cohort of 29 individuals all affected with the same new genetic disorder.”
It is likely to be much more widespread, but still comparatively very rare, she says.
“We tend to only find patients in countries with good access to genetic testing—whether diagnostic or research-based—so there will definitely be affected people in other countries, but without the testing to generate the information about this gene, we can’t be linked up. There could be 100 people affected for example, so still very rare.”
There are 14 different genes in human DNA that make this affected part of the spool and researchers have so far found mutations in six of them. Further research using zebrafish confirmed the mutation.
“Normally, if you have that many genes for the same machinery, you would expect one of the other genes to rescue any problems, but this doesn’t seem to be happening here, which is really intriguing as a geneticist.”
The finding is significant, Dr. Bicknell says.
“A very important outcome of this work is to be able to provide answers and information for the families with an affected family member.
“In addition to giving them an answer as to the cause of the condition, because of the large number of people we have studied, we can provide information to help with managing the disorder, and what might happen in the future. For example, we have several adults affected with the disorder, so this means we are able to give that information to families with young children, that you can expect your child to survive to adulthood.”

From a biology perspective, it gives greater insight into the factors required for brain function. It also suggests that other genetic changes in the same genes might contribute to other disorders affecting brain functioning, that are more complex in origin, such as autism, she says.
“We are seeking to better understand the exact mechanisms, using brain cells grown in a dish, to understand why brain cell functioning is not operating as expected. In the longer term, we want to take advantage of this intriguing observation about the multiple genes involved—can we use this to our advantage in thinking about therapeutic strategies?
“As always, our insights bring about even more questions, so we are starting to follow some of these up in the lab, and in collaboration with other groups,” Dr. Bicknell says.
“We are also helping all of the families involved start a Facebook group, so we can learn from each other and support each other as much as possible.”
About this genetics research news
Author: Press Office
Source: University of Otago
Contact: Press Office – University of Otago
Image: The image is in the public domain
Original Research: Open access.
“Recurrent de novo missense variants across multiple histone H4 genes underlie a neurodevelopmental syndrome” by Federico Tessadori et al. American Journal of Human Genetics
Abstract
Recurrent de novo missense variants across multiple histone H4 genes underlie a neurodevelopmental syndrome
Chromatin is essentially an array of nucleosomes, each of which consists of the DNA double-stranded fiber wrapped around a histone octamer. This organization supports cellular processes such as DNA replication, DNA transcription, and DNA repair in all eukaryotes.
Human histone H4 is encoded by fourteen canonical histone H4 genes, all differing at the nucleotide level but encoding an invariant protein. Here, we present a cohort of 29 subjects with de novo missense variants in six H4 genes (H4C3, H4C4, H4C5, H4C6, H4C9, and H4C11) identified by whole-exome sequencing and matchmaking. All individuals present with neurodevelopmental features of intellectual disability and motor and/or gross developmental delay, while non-neurological features are more variable.
Ten amino acids are affected, six recurrently, and are all located within the H4 core or C-terminal tail. These variants cluster to specific regions of the core H4 globular domain, where protein-protein interactions occur with either other histone subunits or histone chaperones. Functional consequences of the identified variants were evaluated in zebrafish embryos, which displayed abnormal general development, defective head organs, and reduced body axis length, providing compelling evidence for the causality of the reported disorder(s).
While multiple developmental syndromes have been linked to chromatin-associated factors, missense-bearing histone variants (e.g., H3 oncohistones) are only recently emerging as a major cause of pathogenicity. Our findings establish a broader involvement of H4 variants in developmental syndromes.