Howard Hughes Medical Institute (HHMI) scientists have developed a strategy for finding disease-causing mutations that lurk in only a small fraction of the body’s cells. Such mutations can cause significant problems, but cannot be detected with traditional methods of genetic testing, as well as newer, more costly genome sequencing technologies.
The scientists report in the August 21, 2014, issue of the New England Journal of Medicine, that they used the technique to find disease-causing mutations in patients with brain malformations whose genetic causes were unknown despite previous testing.
By sequencing hundreds of copies of the genes in a panel of candidate genes, scientists led by HHMI investigator Christopher A. Walsh identified somatic mutations—gene mutations present in some, but nor all, cells – in more than a quarter of patients that could be successfully diagnosed genetically. Walsh and his colleagues, including Saumya Jamuar, a clinical fellow in Walsh’s lab at Boston Children’s Hospital who is now at the KK Women’s and Children’s Hospital in Singapore and Timothy Yu, also at Boston Children’s Hospital, were authors of the study.
Walsh says his team was surprised to discover so many somatic mutations in patients who had already undergone genetic testing. “This tells us just how poorly other methods perform in detecting somatic mutations,” he says. “You’re not going to find these things unless you go looking for them—unless you have a clinical test that is set up to detect them in a sensitive way.”
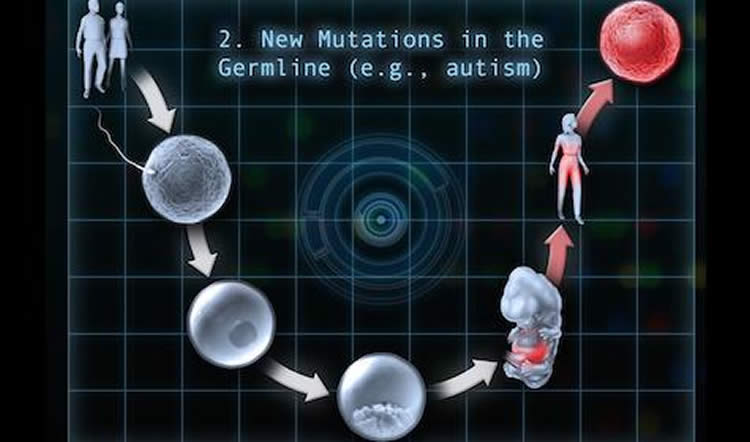
Somatic mutations are not inherited from parents, but instead, arise sometime after fertilization. They are most often seen in some forms of cancer, in which genetic differences between tumor cells and the rest of the body drive tumor growth and metastasis. But they’ve been implicated in other diseases, as well.
“Somatic mutations have been discovered to cause milder forms of a wide range of diseases, especially neuropsychiatric ones,” Walsh says, citing as examples Rett syndrome and tuberous sclerosis, two disorders that cause seizures and intellectual disability. In his own lab, he had found that some of his patients with double cortex syndrome, a brain malformation that can cause some of the same kinds of neurological problems, have somatic mutations. And in a new study published August 21, 2014, in Cell Reports, Walsh’s team analyzed the genomes of individual cells in healthy and diseased brains and found that large segments of DNA had been duplicated or deleted in most cells. “No neuron’s genome is pristine,” he says. “There’s a lot of variability, and some of these mutations have occurred at a stage where they’re present in more than one cell.”
“We think these somatic mutations are probably more common as causes of intellectual disability, and maybe even some psychiatric conditions, than people have generally realized.” Walsh says. “It’s really time to start investigating that systematically.”
He decided to begin with his own patients. A genetic diagnosis is important for counseling patients and their parents about risks to future children, and can, in some cases, influence treatment decisions. But many patients who had come to Walsh’s lab with neurodevelopmental problems were still without answers. “We’d successfully identified causative mutations in many families. But there remained a subset where—even after 10 or more years of searching—we had been unable to identify the causative genes. This made us wonder whether there might be certain kinds of mutations not well discovered by present methods,” he says.
Walsh’s team questioned whether it had missed somatic mutations in those patients by using traditional methods of genetic testing? It seemed possible. Those techniques are not designed to find mutations that occur only in a small fraction of cells, Walsh says. “Even if you are looking at the right gene, you can still miss the mutation.”
Most diagnostic gene testing is done by sequencing specific genes using a traditional DNA sequencing technique known as the Sanger method. When this strategy fails, the search for mutations is sometimes broadened to all of the protein-coding regions of the genome—the exome—or further, to the entire genome. Both approaches have limitations, Walsh says.
“Whole-exome sequencing tends to sample the genome about 30 or 50 times over,” he explains. “But if a mutation is only in five or 10 percent of the cells, then it’s only going to be in a very small fraction of the data, and it’s hard to separate from the noise. The same is true of Sanger sequencing: it has not been optimized to detect a mutation that’s present in a small fraction of the reads.”
To find the kinds of mutations they were looking for, Walsh’s team knew they would have to deepen their search. They devised a strategy in which they would use next-generation sequencing technology to sequence a panel of genes known or suspected to be associated with brain malformations. “We said we’d shoot to sequence them a thousand times over,” Walsh says. “Even if a mutation is only present in five percent of the cells, it will be obvious that it’s a mutation, because we’ll see that mutation 50 times.”
Jamuar set up a test to screen blood samples from 158 patients whose brain malformations remained unexplained. For each patient, a panel of 14 or 54 genes (depending on the patient’s condition) was sequenced hundreds or thousands of times. The design of the panel and sequencing took about 2-3 months to carry out. He then fine-tuned existing bioinformatics algorithms to search for somatic mutations in the sequences. Though the initial sequencing was fast, follow-up validation of potential somatic mutations took additional months because it remains labor-intensive.
In this way, the team uncovered mutations likely to cause disease—either because their role was already known or because they disrupted protein function—in 27 of the 158 patients in their study. Eight of these were somatic mutations, present in just five to 35 percent of the sequenced DNA. Jamuar confirmed these sequencing findings with laboratory experiments in which the patients’ DNA was replicated in bacterial cells and analyzed by Sanger sequencing.
“We have a genetic diagnosis. This ends the diagnostic odyssey for these eight individuals,” says Jamuar.
Five of the eight somatic mutations that they identified would never have been found with traditional sequencing methods, the scientists say. “All of the mutations that were present at less than about 15 percent of the reads were completely undetectable by Sanger sequencing,” Walsh says. “One of them had been missed by whole-exome sequencing, as well.”
“The gold standard of clinical diagnosis is Sanger sequencing,” Jamuar adds. “But you’re missing a big chunk of patients with mutations in these genes, because you are using a test that’s not designed to look for them.”
Now that they have demonstrated their method’s sensitivity in detecting somatic mutations, Walsh and Jamuar say medical geneticists should consider using the approach before turning to more costly whole-exome sequencing. Neither offers a single solution for all patients, but their complementary strengths give geneticists a more complete set of tools. “Look deep, and you may find the answer,” says Jamuar.
Source: Jim Keeley – Howard Hughes Medical Institute
Contact: Howard Hughes Medical Institute press release
Image Source: The image is credited to HHMI BioInteractive and is adapted from the press release
Video Source: The video is available at the Howard Hughes Medical Institute website
Original Research: Abstract for “Somatic Mutations in Cerebral Cortical Malformations” by Saumya S. Jamuar, M.R.C.P.C.H., Anh-Thu N. Lam, B.S., Martin Kircher, Ph.D., Alissa M. D’Gama, B.A., Jian Wang, Ph.D., Brenda J. Barry, M.S., Xiaochang Zhang, Ph.D., Robert Sean Hill, Ph.D., Jennifer N. Partlow, M.S., Aldo Rozzo, D.V.M., Ph.D., Sarah Servattalab, B.S., Bhaven K. Mehta, M.A., Meral Topcu, Ph.D., Dina Amrom, M.D., Eva Andermann, M.D., Ph.D., Bernard Dan, Ph.D., Elena Parrini, Ph.D., Renzo Guerrini, M.D., Ingrid E. Scheffer, M.B., B.S., Ph.D., Samuel F. Berkovic, M.D., Richard J. Leventer, M.B., B.S., Ph.D., Yiping Shen, Ph.D., Bai Lin Wu, Ph.D., A. James Barkovich, M.D., Mustafa Sahin, M.D., Ph.D., Bernard S. Chang, M.D., Michael Bamshad, M.D., Deborah A. Nickerson, Ph.D., Jay Shendure, M.D., Ph.D., Annapurna Poduri, M.D., M.P.H., Timothy W. Yu, M.D., Ph.D., and Christopher A. Walsh, M.D., Ph.D. in New England Journal of Medicine. Published online August 21 2014 doi:10.1056/NEJMoa1314432
Full open access research for “CC2D1A Regulates Human Intellectual and Social Function as well as NF-κB Signaling Homeostasis” by M. Chiara Manzini, Lan Xiong, Ranad Shaheen, Dimira E. Tambunan, Stefania Di Costanzo, Vanessa Mitisalis, David J. Tischfield, Antonella Cinquino, Mohammed Ghaziuddin, Mehtab Christian, Qin Jiang, Sandra Laurent, Zohair A. Nanjiani, Saima Rasheed, R. Sean Hill, Sofia B. Lizarraga, Danielle Gleason, Diya Sabbagh, Mustafa A. Salih, Fowzan S. Alkuraya, Christopher A. Walsh in Cell Reports. Published online August 2014 doi:10.1016/j.celrep.2014.06.039
CC2D1A Regulates Human Intellectual and Social Function as well as NF-κB Signaling Homeostasis
Autism spectrum disorder (ASD) and intellectual disability (ID) are often comorbid, but the extent to which they share common genetic causes remains controversial. Here, we present two autosomal-recessive “founder” mutations in the CC2D1A gene causing fully penetrant cognitive phenotypes, including mild-to-severe ID, ASD, as well as seizures, suggesting shared developmental mechanisms. CC2D1A regulates multiple intracellular signaling pathways, and we found its strongest effect to be on the transcription factor nuclear factor κB (NF-κB). Cc2d1a gain and loss of function both increase activation of NF-κB, revealing a critical role of Cc2d1a in homeostatic control of intracellular signaling. Cc2d1a knockdown in neurons reduces dendritic complexity and increases NF-κB activity, and the effects of Cc2d1a depletion can be rescued by inhibiting NF-κB activity. Homeostatic regulation of neuronal signaling pathways provides a mechanism whereby common founder mutations could manifest diverse symptoms in different patients.
“CC2D1A Regulates Human Intellectual and Social Function as well as NF-κB Signaling Homeostasis” by M. Chiara Manzini, Lan Xiong, Ranad Shaheen, Dimira E. Tambunan, Stefania Di Costanzo, Vanessa Mitisalis, David J. Tischfield, Antonella Cinquino, Mohammed Ghaziuddin, Mehtab Christian, Qin Jiang, Sandra Laurent, Zohair A. Nanjiani, Saima Rasheed, R. Sean Hill, Sofia B. Lizarraga, Danielle Gleason, Diya Sabbagh, Mustafa A. Salih, Fowzan S. Alkuraya, Christopher A. Walsh in Cell Reports, August 2014 doi:10.1016/j.celrep.2014.06.039