Summary: The adult nervous system of a C. elegans worm contains 302 neurons, which can be divided into 118 types. Seventy homeobox genes are involved in characterizing the neuron types.
Source: NIH/NINDS
When it comes to brain cells, one size does not fit all. Neurons come in a wide variety of shapes, sizes, and contain different types of brain chemicals. But how did they get that way? A new study in Nature suggests that the identities of all the neurons in a worm are linked to unique members of a single gene family that control the process of converting DNA instructions into proteins, known as gene expression. The results of this study could provide a foundation for understanding how nervous systems have evolved in many other animals, including humans. The study was funded by the National Institute of Neurological Disorders and Stroke (NINDS), part of the National Institutes of Health.
“The central nervous systems of all animals, from worms to humans, are incredibly intricate and highly ordered. The generation and diversity of a plethora of neuronal cell types is driven by gene expression,” said Robert Riddle, Ph.D., program director at NINDS. “So, it is surprising and exciting to consider that the cell diversity we see in the entire nervous system could come from a just a single group of genes.”
Researchers led by Oliver Hobert, Ph.D., professor of biochemistry and molecular biophysics at Columbia University in New York City and graduate student Molly B. Reilly, wanted to know how brain cells in the C. elegans worm got their various shapes and functions. For these experiments, the researchers used a genetically engineered worm in which individual neurons were color coded. In addition, coding sequences for green fluorescence protein were inserted into homeobox genes, a highly conserved set of genes known to play fundamental roles in development. Homeobox gene expression patterns were determined by examining the patterns of the glowing fluorescent marker.
Dr. Hobert’s team discovered that across the entire worm nervous system, each type of neuron contained a unique set of homeobox proteins. In other words, the identity of every neuron could be traced to a specific combination of homeobox genes that were switched on or off.
Homeobox genes were initially discovered for their role in ensuring that body parts end up in their proper places. These genes have since been found across all animal species, as well as in plants and fungi. Homeobox genes contain instructions for making transcription factors, which are proteins that can control the activity of other genes.
The nervous system of an adult C. elegans contains 302 neurons, which can be divided into 118 types. Dr. Hobert’s team determined that 70 homeobox genes are involved in characterizing the neuronal types. In addition, further analysis suggested that the homeobox codes could be used to subdivide classes of neurons. For example, the additional analysis identified not only that a cell was a motor neuron, but its location in the ventral nerve cord, the worm’s version of a spinal cord.
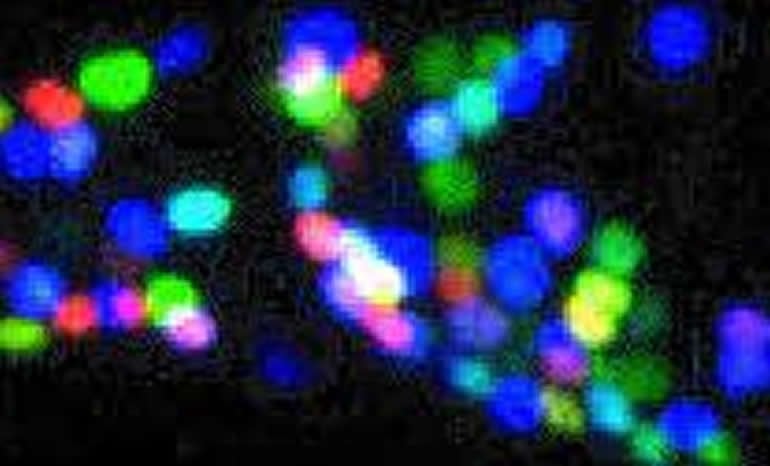
“The diversity in neurons may be driven by a gene family that distinguishes individual cell types in the brain,” said Dr. Hobert, “There is a simplicity to this arrangement in the sense that just one gene family may explain all brain cell types. The complexity that arose from this simple ‘barcode’ gene family suggests a potential role for homeobox genes in the evolution of the nervous system.”
Future studies will investigate whether brain cells in other organisms can also be identified by specific protein codes. In addition, researchers will examine the effects of changing the codes on identity and function of neurons.
Funding: This study was supported by NINDS (F31 NS105398, R21 NS106843).
About this genetics research article
Source:
NIH/NINDS
Contacts:
Barbara McMakin – NIH/NINDS
Image Source:
The image is credited to Hobert Lab, Columbia University.
Original Research: Closed access
“Unique homeobox codes delineate all C. elegans neuron classes” by Molly B. Reilly, Cyril Cros, Erdem Varol, Eviatar Yemini & Oliver Hobert. Nature.
Abstract
Unique homeobox codes delineate all C. elegans neuron classes
It is not known at present whether neuronal cell-type diversity—defined by cell-type-specific anatomical, biophysical, functional and molecular signatures—can be reduced to relatively simple molecular descriptors of neuronal identity1. Here we show, through examination of the expression of all of the conserved homeodomain proteins encoded by the Caenorhabditis elegans genome2, that the complete set of 118 neuron classes of C. elegans can be described individually by unique combinations of the expression of homeodomain proteins, thereby providing—to our knowledge—the simplest currently known descriptor of neuronal diversity. Computational and genetic loss-of-function analyses corroborate the notion that homeodomain proteins not only provide unique descriptors of neuron type, but also have a critical role in specifying neuronal identity. We speculate that the pervasive use of homeobox genes in defining unique neuronal identities reflects the evolutionary history of neuronal cell-type specification.