Summary: Study identifies thirty-two gene regions, including nine novel gene regions, associated with facial features including the shape of the face, lips, and nose.
Source: UCL
Genes that determine the shape of a person’s facial profile have been discovered by a UCL-led research team.
The researchers identified 32 gene regions that influenced facial features such as nose, lip, jaw, and brow shape, nine of which were entirely new discoveries while the others validated genes with prior limited evidence.
The analysis of data from more than 6,000 volunteers across Latin America was published today in Science Advances.
The international research team, led from UCL, Aix-Marseille University and The Open University, found that one of the genes appears to have been inherited from the Denisovans, an extinct group of ancient humans who lived tens of thousands of years ago.
The team found that the gene, TBX15, which contributes to lip shape, was linked with genetic data found in the Denisovan people, providing a clue to the gene’s origin. The Denisovans lived in central Asia, and other studies suggest they interbred with modern humans, as some of their DNA lives on in Pacific Islanders and Indigenous people of the Americas.
Co-corresponding author Dr Kaustubh Adhikari (UCL Genetics, Evolution & Environment and The Open University) said: “The face shape genes we found may have been the product of evolution as ancient humans evolved to adapt to their environments. Possibly, the version of the gene determining lip shape that was present in the Denisovans could have helped in body fat distribution to make them better suited to the cold climates of Central Asia, and was passed on to modern humans when the two groups met and interbred.”
Co-first author Dr Pierre Faux (Aix-Marseille University) said: “To our knowledge this is the first time that a version of a gene inherited from ancient humans is associated with a facial feature in modern humans. In this case, it was only possible because we moved beyond Eurocentric research; modern-day Europeans do not carry any DNA from the Denisovans, but Native Americans do.”
Co-first author Betty Bonfante (Aix-Marseille University) added: “It is one of only a few studies looking for genes affecting the face in a non-European population, and the first one to focus on the profile only.”
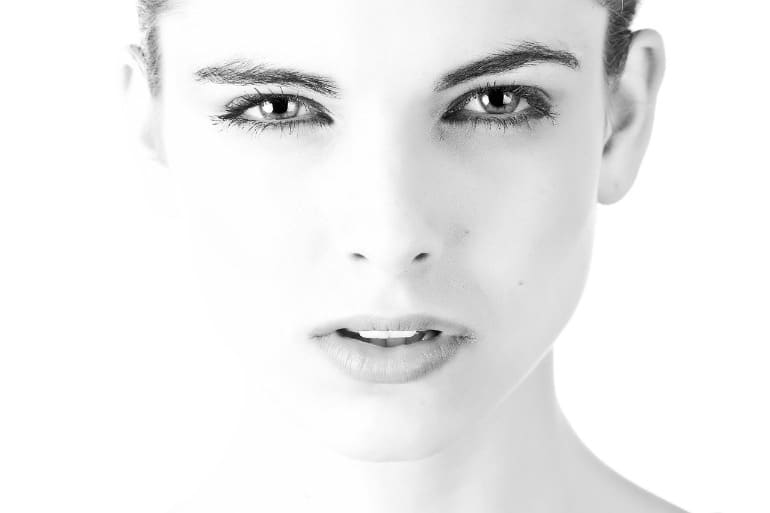

Researchers have only been able to analyse complex genetic data from thousands of people at once over the last two decades, since the mapping of the human genome enabled the use of genome-wide association studies to find correlations between traits and genes. This study compared genetic information from the study participants with characteristics of their face shape, quantified with 59 measurements (distances, angles and ratios between set points) from photos of the participants’ faces in profile.
Co-corresponding author Professor Andres Ruiz-Linares (Fudan University, UCL Genetics, Evolution & Environment, and Aix-Marseille University) said: “Research like this can provide basic biomedical insights and help us understand how humans evolved.”
The findings of this research could help understand the developmental processes that determine facial features, which will help researchers studying genetic disorders that lead to facial abnormalities.
The results also contribute to the understanding of the evolution of facial appearance in human and other species. One of the newly discovered genes found in this study is VPS13B, which influenced nose pointiness; the researchers also found that this gene affects nose structure in mice, indicating a broadly shared genetic basis among distantly related mammal species.
About this genetics research news
Source: UCL
Contact: Chris Lane – UCL
Image: The image is in the public domain
Original Research: Open access.
“A GWAS in Latin Americans identifies novel face shape loci, implicating VPS13B and a Denisovan introgressed region in facial variation” by Betty Bonfante et al. Science Advances
Abstract
A GWAS in Latin Americans identifies novel face shape loci, implicating VPS13B and a Denisovan introgressed region in facial variation
To characterize the genetic basis of facial features in Latin Americans, we performed a genome-wide association study (GWAS) of more than 6000 individuals using 59 landmark-based measurements from two-dimensional profile photographs and ~9,000,000 genotyped or imputed single-nucleotide polymorphisms. We detected significant association of 32 traits with at least 1 (and up to 6) of 32 different genomic regions, more than doubling the number of robustly associated face morphology loci reported until now (from 11 to 23). These GWAS hits are strongly enriched in regulatory sequences active specifically during craniofacial development. The associated region in 1p12 includes a tract of archaic adaptive introgression, with a Denisovan haplotype common in Native Americans affecting particularly lip thickness. Among the nine previously unidentified face morphology loci we identified is the VPS13B gene region, and we show that variants in this region also affect midfacial morphology in mice.