Summary: Using CRISPR-Cas9 gene editing, researchers identified actionable pathways responsible for the growth of glioblastoma stem cells. By reverse engineering brain cancer cells, multiple potential new targets for cancer treatments have been uncovered.
Source: University of Toronto
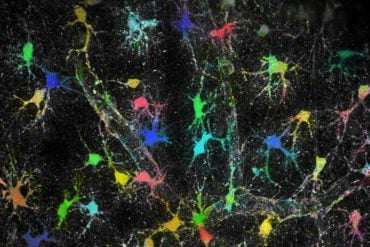
Glioblastoma is one of the most devastating forms of cancer, with few existing treatment options. It is also a leading cause of cancer-related death in children and young adults. Scientists have ‘reverse engineered’ brain cancer stem cells gene by gene, uncovering multiple potential targets for this hard-to-treat cancer.
This work is a collaboration between the University of Toronto, The Hospital for Sick Children (SickKids), and the University of Calgary. Findings were published today in the journal Cell Reports, making this the first published study to systematically profile a large panel of patient-derived brain tumour cells that have stem cell properties.
“We think that, in one big experiment, we have uncovered many new targets for glioblastoma, some of which were surprising,” says Dr. Peter Dirks, co-principal investigator of the study, Staff Neurosurgeon and Senior Scientist at SickKids. “These glioblastoma stem cells are also resistant to treatment, which is one reason that these tumours are so hard to cure. We need new ways to disrupt these cells specifically if we are going to give people a better chance of survival.”
The research team also found that adult glioblastoma cells are actually dependent on the same genes that are important for brain development in infancy and early childhood. “This really emphasizes how much research needs to be done to understand the developing human brain,” says Dirks, who, in 2003, was the first to discover the existence of cancer stem cells in brain tumours.
CRISPR: A powerful new tool
The emergence of the CRISPR-Cas9 technology provides a powerful new way to explore cancer biology through genome-wide screens. Dr. Stéphane Angers, co-principal investigator of the study and Professor at the Leslie Dan Faculty of Pharmacy, University of Toronto, specializes in the use of CRISPR-Cas9 in cancer. Taking 10 unique patient-derived glioblastoma stem cell cultures collected by the Dirks research team, the Angers lab used CRISPR ‘cell fitness screens’ to determine which genes in the cancer stem cells were required for the cells to survive and to grow, therefore, important for tumour progression.
“Cancer stem cells fuel the growth of tumours and progression of the disease,” says Angers. “In order to effectively target these cells, having a comprehensive view of the genes controlling the growth programs is critical. If you know which genes are necessary for these cells to survive and proliferate, you can then look at ways to attack or block these genes and stop tumour growth in its tracks.”
By systematically knocking out each of the 20,000 genes, one at the time, from each of the 10 patient samples, the Angers team found multiple genetic vulnerabilities and revealed a wealth of data that can be further mined to identify possible drug targets for glioblastoma. “This is one of the first studies of its kind, where CRISPR screens are performed directly in multiple freshly isolated patient cells in parallel. This study has provided a massive amount of new information that the research community can now interrogate to help design new treatment strategies. ” says Angers.
One gene identified in the study, known as DOT1L, was found to be necessary for tumour persistence in seven of the 10 glioblastoma patient tumour cultures. In collaboration with Dr. Samuel Weiss at the Cumming School of Medicine, University of Calgary, the team used preclinical models to demonstrate the effectiveness of a drug currently used to treat leukemia to inhibit the DOT1L gene product in glioblastoma stem cells.
“We found that blocking this specific protein in this particular form of brain cancer reduced tumour growth and resulted in longer survival in the preclinical model,” says Angers.
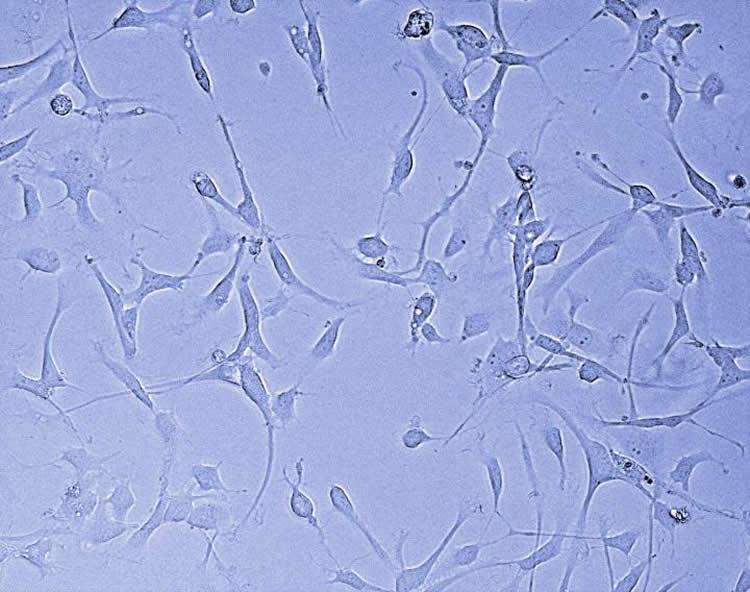
“This is promising because it uncovered a biological process, not previously suspected to be implicated in glioblastoma, for which a small molecule drug already exists.”
Moving beyond a “static picture” of cancer
In recent years, significant time, effort and funding dollars have been spent on the genomic sequencing of cancer tumours. While this has given us a clearer picture of the hundreds of genetic mutations present in glioblastoma and other cancers, for glioblastoma it has not led to any significant treatment advances, Angers says.
“This shows that just knowing about genetic mutations is not enough,” says Graham MacLeod, a post-doctoral fellow with the Angers lab and co-first author of the study. “That is a static picture of cancer. We are learning that we need to better understand the blueprint of how this cancer functions and what specific genes fuel tumour growth in order to attack it.”
Funding: This research was supported by Stand Up to Cancer Canada Cancer Stem Cell Dream Team funding provided by the Government of Canada through Genome Canada and the Canadian Institutes of Health Research, the Ontario Institute for Cancer Research, and SickKids Foundation.
Source:
University of Toronto
Media Contacts:
Kate Richards – University of Toronto
Image Source:
The image is credited to Angers Lab, Leslie Dan Faculty of Pharmacy, University of Toronto.
Original Research: Open access.
“Genome-Wide CRISPR-Cas9 Screens Expose Genetic Vulnerabilities and Mechanisms of Temozolomide Sensitivity in Glioblastoma Stem Cells”. Peter B. Dirks et al. Cell Reports. doi:10.1016/j.celrep.2019.03.047
Abstract
Genome-Wide CRISPR-Cas9 Screens Expose Genetic Vulnerabilities and Mechanisms of Temozolomide Sensitivity in Glioblastoma Stem Cells
Highlights
• Genome-wide CRISPR-Cas9 screens in patient-derived glioblastoma stem cells
• Identification of regulators of stemness governing glioblastoma stem cell growth
• Multiple stress response pathways are genetic vulnerabilities in glioblastoma
• Identification of modulators of sensitivity to standard of care chemotherapy
Summary
Glioblastoma therapies have remained elusive due to limitations in understanding mechanisms of growth and survival of the tumorigenic population. Using CRISPR-Cas9 approaches in patient-derived GBM stem cells (GSCs) to interrogate function of the coding genome, we identify actionable pathways responsible for growth, which reveal the gene-essential circuitry of GBM stemness and proliferation. In particular, we characterize members of the SOX transcription factor family, SOCS3, USP8, and DOT1L, and protein ufmylation as important for GSC growth. Additionally, we reveal mechanisms of temozolomide resistance that could lead to combination strategies. By reaching beyond static genome analysis of bulk tumors, with a genome-wide functional approach, we reveal genetic dependencies within a broad range of biological processes to provide increased understanding of GBM growth and treatment resistance.