Summary: Certain progenitor cells in the bronchi are mainly responsible for producing coronavirus receptors. These cells normally develop into respiratory tract sells linked with cilia, that clear bacteria out of the lungs.
Source: Berlin Institute of Health
Scientists from the Berlin Institute of Health (BIH), Charité – Universitätsmedizin Berlin and the Thorax Clinic at Heidelberg University Hospital, whose collaboration is taking place under the auspices of the German Center for Lung Research (DZL), have examined samples from non-virus infected patients to determine which cells of the lungs and bronchi are targets for novel coronavirus (SARS-CoV-2) infection. They discovered that the receptor for this coronavirus is abundantly expressed in certain progenitor cells. These cells normally develop into respiratory tract cells lined with hair-like projections called cilia that sweep mucus and bacteria out of the lungs. The scientists have now published their findings in The EMBO Journal.
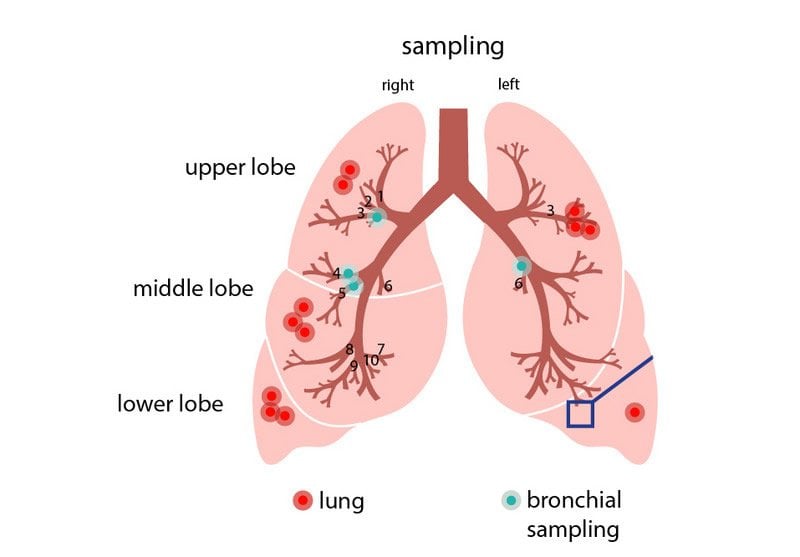
Professor Roland Eils and his colleagues from the Thorax Clinic in Heidelberg initially intended to study why lung cancer sometimes occurs in people who have never smoked. They began by analyzing samples of twelve lung cancer patients. These samples were obtained from the Heidelberg Lung Biobank and came from both the cancerous part of the lungs and the surrounding healthy lung tissue. They also studied cells from the airways of healthy patients, which had been collected in a minimally invasive manner during a bronchoscopy examination performed to rule out lung cancer. The rapidly spreading coronavirus prompted the researchers to take another look at these existing but so far unpublished data. “I was convinced that the data we gathered from these non-coronavirus infected patients would provide important information for understanding the viral infection,” says Roland Eils, founding director of the BIH Digital Health Center.
Infection requires receptors and cofactors
“We wanted to find out which specific cells the coronavirus attacks,” explains Professor Christian Conrad, who also works at the BIH Digital Health Center. The scientists knew, from studies by BIH Professor Christian Drosten, director of the Institute of Virology at Campus Charité; Mitte, and by others, that the virus’s spike protein attaches to an ACE2 receptor on the cell surface. In addition, the virus needs one or more cofactors for it to be able to penetrate cells. But which cells are endowed with such receptors and cofactors? Which cells in which part of the respiratory system are particularly susceptible to SARS-CoV-2 infection? Eils and his colleagues at the BIH and Charité; now used single-cell sequencing technology to examine the cells in the samples from Heidelberg.
60,000 single cells were sequenced
“We then analyzed a total of nearly 60,000 cells to determine whether they activated the gene for the receptor and potential cofactors, thus in principle allowing them to be infected by the coronavirus,” reports Soeren Lukassen, one of the lead authors of the study now being published in The EMBO Journal. “We only found the gene transcripts for ACE2 and for the cofactor TMPRSS2 in very few cells, and only in very small numbers.” Lukassen and his four co-lead authors Robert Lorenz Chua, Timo Trefzer, Nicolas C. Kahn and Marc A. Schneider discovered that certain progenitor cells in the bronchi are mainly responsible for producing the coronavirus receptors. These progenitor cells normally develop into respiratory tract cells lined with hair-like projections called cilia that sweep mucus and bacteria out of the lungs. “Armed with the knowledge of which cells are attacked, we can now develop targeted therapies,” explains Professor Michael Kreuter from the Thorax Clinic at Heidelberg University Hospital.
Why does the infection progress so differently?
An interesting additional finding of the study was that the ACE2 receptor density on the cells increased with age and was generally higher in men than in women. “This was only a trend, but it could explain why SARS-CoV-2 has infected more men than women,” Eils says. However, he points out, “our sample sizes are still much too small to make conclusive statements, so we need to repeat the study in larger patient cohorts.”
“These results show us that the virus acts in a highly selective manner, and that it is dependent on certain human cells in order to spread and replicate,” Eils explains. “The better we understand the interaction between the virus and its host, the better we will be able to develop effective counterstrategies.” He and the other researchers will next study COVID 19 patients to ascertain whether the virus has actually infected these cells. “We want to understand why the infection takes a benign course in some patients, while causing severe disease in others,” Eils says. “So we will also look closely at the immune cells in the infected tissue.”
Tech firms provide expertise
Based on a high-performance technology solution designed by Intel, Dell developed a hardware and system architecture that reduced the processing time needed to sequence the 60,000 single cells. Hannes Schwaderer, country manager of Intel Germany, explains: “There are many things we don’t know about the coronavirus. This research project and the next steps require enormous computing resources. That’s exactly where our expertise can help.”
BIH is supporting COVID-19 research
The Berlin Institute of Health (BIH) is helping advance research into the novel coronavirus, SARS-CoV-2, through a targeted program. Professor Axel R. Pries, interim Chairman of the BIH Executive Board and Dean of Charité – Universitätsmedizin, stresses: “In light of the global threat posed by the SARS-CoV-2 virus, we as researchers have a duty to marshal all of our scientific knowledge to understand the virus and its infection strategies as well as the disease progression of COVID-19 patients. Only so can we better identify high-risk patients and develop new therapies and vaccines. Every contribution towards this effort makes a difference.”
About this coronavirus research article
Source:
Berlin Institute of Health
Media Contacts:
Stefanie Seltmann – Berlin Institute of Health
Image Source:
The image is credited to the researchers.
Original Research: Closed access
“SARS‐CoV‐2 receptor ACE2 and TMPRSS2 are primarily expressed in bronchial transient secretory cells”. Soeren Lukassen, Robert Lorenz Chua, Timo Trefzer, Nicolas C. Kahn, Marc A. Schneider, Thomas Muley, Hauke Winter, Michael Meister, Carmen Veith, Agnes W. Boots, Bianca P. Hennig, Michael Kreuter, Christian Conrad & Roland Eils.
EMBO Journal doi:10.15252/embj.20105114.
Abstract
SARS‐CoV‐2 receptor ACE2 and TMPRSS2 are primarily expressed in bronchial transient secretory cells
The SARS‐CoV‐2 pandemic affecting the human respiratory system severely challenges public health and urgently demands for increasing our understanding of COVID‐19 pathogenesis, especially host factors facilitating virus infection and replication. SARS‐CoV‐2 was reported to enter cells via binding to ACE2, followed by its priming by TMPRSS2. Here, we investigate ACE2 and TMPRSS2 expression levels and their distribution across cell types in lung tissue (twelve donors, 39,778 cells) and in cells derived from subsegmental bronchial branches (four donors, 17,521 cells) by single nuclei and single cell RNA sequencing, respectively. While TMPRSS2 is expressed in both tissues, in the subsegmental bronchial branches ACE2 is predominantly expressed in a transient secretory cell type. Interestingly, these transiently differentiating cells show an enrichment for pathways related to RHO GTPase function and viral processes suggesting increased vulnerability for SARS‐CoV‐2 infection. Our data provide a rich resource for future investigations of COVID‐19 infection and pathogenesis.
Feel Free To Share This COVID-19 News.