Summary: The newly developed HeadXNet deep learning system can accurately detect clinically significant aneurysms from patients’ brain scans. The tool could improve diagnostics and care for those experiencing aneurysms.
Source: Stanford
Doctors could soon get some help from an artificial intelligence tool when diagnosing brain aneurysms – bulges in blood vessels in the brain that can leak or burst open, potentially leading to stroke, brain damage or death.
The AI tool, developed by researchers at Stanford University and detailed in a paper published June 7 in JAMA Network Open, highlights areas of a brain scan that are likely to contain an aneurysm.
“There’s been a lot of concern about how machine learning will actually work within the medical field,” said Allison Park, a Stanford graduate student in statistics and co-lead author of the paper. “This research is an example of how humans stay involved in the diagnostic process, aided by an artificial intelligence tool.”
This tool, which is built around an algorithm called HeadXNet, improved clinicians’ ability to correctly identify aneurysms at a level equivalent to finding six more aneurysms in 100 scans that contain aneurysms. It also improved consensus among the interpreting clinicians. While the success of HeadXNet in these experiments is promising, the team of researchers – who have expertise in machine learning, radiology and neurosurgery – cautions that further investigation is needed to evaluate the generalizability of the AI tool prior to real-time clinical deployment given differences in scanner hardware and imaging protocols across different hospital centers. The researchers plan to address such problems through multi-center collaboration.
Augmented expertise
Combing brain scans for signs of an aneurysm can mean scrolling through hundreds of images. Aneurysms come in many sizes and shapes and balloon out at tricky angles – some register as no more than a blip within the movie-like succession of images.
“Search for an aneurysm is one of the most labor-intensive and critical tasks radiologists undertake,” said Kristen Yeom, associate professor of radiology and co-senior author of the paper. “Given inherent challenges of complex neurovascular anatomy and potential fatal outcome of a missed aneurysm, it prompted me to apply advances in computer science and vision to neuroimaging.”
Yeom brought the idea to the AI for Healthcare Bootcamp run by Stanford’s Machine Learning Group, which is led by Andrew Ng, adjunct professor of computer science and co-senior author of the paper. The central challenge was creating an artificial intelligence tool that could accurately process these large stacks of 3D images and complement clinical diagnostic practice.
To train their algorithm, Yeom worked with Park and Christopher Chute, a graduate student in computer science, and outlined clinically significant aneurysms detectable on 611 computerized tomography (CT) angiogram head scans.
“We labelled, by hand, every voxel – the 3D equivalent to a pixel – with whether or not it was part of an aneurysm,” said Chute, who is also co-lead author of the paper. “Building the training data was a pretty grueling task and there were a lot of data.”
Following the training, the algorithm decides for each voxel of a scan whether there is an aneurysm present. The end result of the HeadXNet tool is the algorithm’s conclusions overlaid as a semi-transparent highlight on top of the scan. This representation of the algorithm’s decision makes it easy for the clinicians to still see what the scans look like without HeadXNet’s input.
“We were interested how these scans with AI-added overlays would improve the performance of clinicians,” said Pranav Rajpurkar, a graduate student in computer science and co-lead author of the paper. “Rather than just having the algorithm say that a scan contained an aneurysm, we were able to bring the exact locations of the aneurysms to the clinician’s attention.”
Eight clinicians tested HeadXNet by evaluating a set of 115 brain scans for aneurysm, once with the help of HeadXNet and once without. With the tool, the clinicians correctly identified more aneurysms, and therefore reduced the “miss” rate, and the clinicians were more likely to agree with one another. HeadXNet did not influence how long it took the clinicians to decide on a diagnosis or their ability to correctly identify scans without aneurysms – a guard against telling someone they have an aneurysm when they don’t.
To other tasks and institutions
The machine learning methods at the heart of HeadXNet could likely be trained to identify other diseases inside and outside the brain. For example, Yeom imagines a future version could focus on speeding up identifying aneurysms after they have burst, saving precious time in an urgent situation. But a considerable hurdle remains in integrating any artificial intelligence medical tools with daily clinical workflow in radiology across hospitals.
Current scan viewers aren’t designed to work with deep learning assistance, so the researchers had to custom-build tools to integrate HeadXNet within scan viewers. Similarly, variations in real-world data – as opposed to the data on which the algorithm is tested and trained – could reduce model performance. If the algorithm processes data from different kinds of scanners or imaging protocols, or a patient population that wasn’t part of its original training, it might not work as expected.
“Because of these issues, I think deployment will come faster not with pure AI automation, but instead with AI and radiologists collaborating,” said Ng. “We still have technical and non-technical work to do, but we as a community will get there and AI-radiologist collaboration is the most promising path.”
Additional Stanford co-authors are Joe Lou, undergraduate in computer science; Robyn Ball, senior biostatistician at the Quantitative Sciences Unit (also affiliated with Roam Analytics); graduate students Katie Shpanskaya, Rashad Jabarkheel, Lily H. Kim and Emily McKenna; radiology residents Joe Tseng and Jason Ni; Fidaa Wishah, clinical instructor of radiology; Fred Wittber, diagnostic radiology fellow; David S. Hong, assistant professor of psychiatry and behavioral sciences; Thomas J. Wilson, clinical assistant professor of neurosurgery; Safwan Halabi, clinical associate professor of radiology; Sanjay Basu, assistant professor of medicine; Bhavik N. Patel, assistant professor of radiology; and Matthew P. Lungren, assistant professor of radiology.
Hong and Yeom are also members of Stanford Bio-X, the Stanford Maternal and Child Health Research Institute and the Wu Tsai Neurosciences Institute. Patel is also a member of Stanford Bio-X and the Stanford Cancer Institute. Lungren is a member of Stanford Bio-X, the Stanford Maternal and Child Health Research Institute and the Stanford Cancer Institute.
Source:
Stanford
Media Contacts:
TTaylor Kubota – Stanford
Image Source:
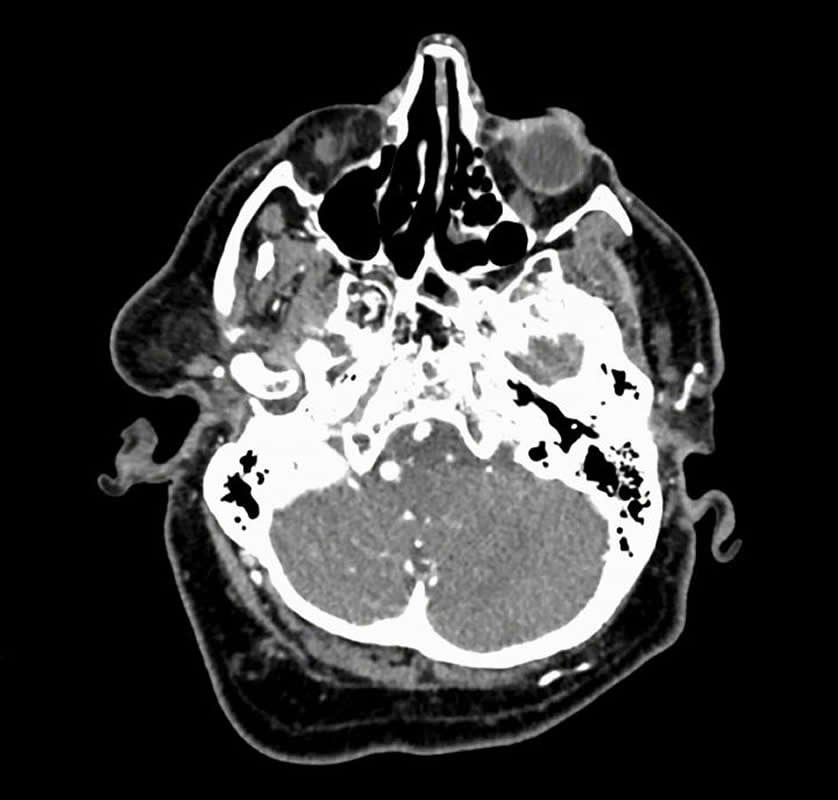
The image is credited to Allison Park.
Original Research: Open access
“Deep Learning–Assisted Diagnosis of Cerebral Aneurysms Using the HeadXNet Model”. Allison Park, BA; Chris Chute, BS; Pranav Rajpurkar, MS; Joe Lou; Robyn L. Ball, PhD; Katie Shpanskaya, BS; Rashad Jabarkheel, BS; Lily H. Kim, BS; Emily McKenna, BS; Joe Tseng, MD; Jason Ni, MD; Fidaa Wishah, MD; Fred Wittber, MD; David S. Hong, MD; Thomas J. Wilson, MD; Safwan Halabi, MD; Sanjay Basu, MD, PhD; Bhavik N. Patel, MD, MBA; Matthew P. Lungren, MD, MPH; Andrew Y. Ng, PhD; Kristen W. Yeom, MD.
JAMA Network Open. doi:10.1001/jamanetworkopen.2019.5600
Abstract
Deep Learning–Assisted Diagnosis of Cerebral Aneurysms Using the HeadXNet Model
Importance Deep learning has the potential to augment clinician performance in medical imaging interpretation and reduce time to diagnosis through automated segmentation. Few studies to date have explored this topic.
Objective To develop and apply a neural network segmentation model (the HeadXNet model) capable of generating precise voxel-by-voxel predictions of intracranial aneurysms on head computed tomographic angiography (CTA) imaging to augment clinicians’ intracranial aneurysm diagnostic performance.
Design, Setting, and Participants In this diagnostic study, a 3-dimensional convolutional neural network architecture was developed using a training set of 611 head CTA examinations to generate aneurysm segmentations. Segmentation outputs from this support model on a test set of 115 examinations were provided to clinicians. Between August 13, 2018, and October 4, 2018, 8 clinicians diagnosed the presence of aneurysm on the test set, both with and without model augmentation, in a crossover design using randomized order and a 14-day washout period. Head and neck examinations performed between January 3, 2003, and May 31, 2017, at a single academic medical center were used to train, validate, and test the model. Examinations positive for aneurysm had at least 1 clinically significant, nonruptured intracranial aneurysm. Examinations with hemorrhage, ruptured aneurysm, posttraumatic or infectious pseudoaneurysm, arteriovenous malformation, surgical clips, coils, catheters, or other surgical hardware were excluded. All other CTA examinations were considered controls.
Main Outcomes and Measures Sensitivity, specificity, accuracy, time, and interrater agreement were measured. Metrics for clinician performance with and without model augmentation were compared.
Results The data set contained 818 examinations from 662 unique patients with 328 CTA examinations (40.1%) containing at least 1 intracranial aneurysm and 490 examinations (59.9%) without intracranial aneurysms. The 8 clinicians reading the test set ranged in experience from 2 to 12 years. Augmenting clinicians with artificial intelligence–produced segmentation predictions resulted in clinicians achieving statistically significant improvements in sensitivity, accuracy, and interrater agreement when compared with no augmentation. The clinicians’ mean sensitivity increased by 0.059 (95% CI, 0.028-0.091; adjusted P = .01), mean accuracy increased by 0.038 (95% CI, 0.014-0.062; adjusted P = .02), and mean interrater agreement (Fleiss κ) increased by 0.060, from 0.799 to 0.859 (adjusted P = .05). There was no statistically significant change in mean specificity (0.016; 95% CI, −0.010 to 0.041; adjusted P = .16) and time to diagnosis (5.71 seconds; 95% CI, 7.22-18.63 seconds; adjusted P = .19).
Conclusions and Relevance The deep learning model developed successfully detected clinically significant intracranial aneurysms on CTA. This suggests that integration of an artificial intelligence–assisted diagnostic model may augment clinician performance with dependable and accurate predictions and thereby optimize patient care.






