In collaboration with several Japanese institutes, researchers at the RIKEN Quantitative Biology Center in Japan demonstrate an easy and fast way to achieve whole brain imaging for 3D analysis of gene expression profiles and neural circuits at the systems level.
*Please see the notes for important information regarding this research.*
A major challenge of systems biology is understanding how phenomena at the cellular scale correlate with activity at the organism level. A concerted effort has been made especially in the brain, as scientists are aiming to clarify how neural activity is translated into consciousness and other complex brain activities. One example of the technologies needed is whole-brain imaging at single-cell resolution. This imaging normally involves preparing a highly transparent sample that minimizes light scattering and then imaging neurons tagged with fluorescent probes at different slices to produce a 3D representation. However, limitations in current methods prevent comprehensive study of the relationship. A new high-throughput method, CUBIC (Clear, Unobstructed Brain Imaging Cocktails and Computational Analysis), published in Cell, is a great leap forward, as it offers unprecedented rapid whole-brain imaging at single cell resolution and a simple protocol to clear and transparentize the brain sample based on the use of aminoalcohols.
In combination with light sheet fluorescence microscopy, CUBIC was tested for rapid imaging of a number of mammalian systems, such as mouse and primate, showing its scalability for brains of different size. Additionally, it was used to acquire new spatial-temporal details of gene expression patterns in the hypothalamic circadian rhythm center. Moreover, by combining images taken from opposite directions, CUBIC enables whole brain imaging and direct comparison of brains in different environmental conditions.
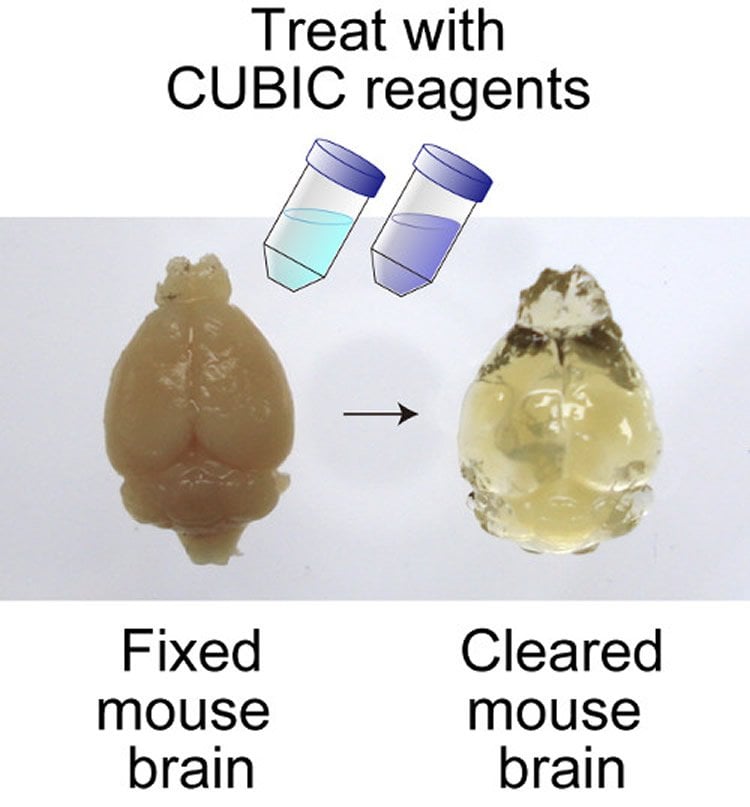
CUBIC overcomes a number of obstacles compared with previous methods. One is the clearing and transparency protocol, which involves serially immersing fixed tissues into just two reagents for a relatively short time. Second, CUBIC is compatible with many fluorescent probes because of low quenching, which allows for probes with longer wavelengths and reduces concern for scattering when whole brain imaging while at the same time inviting multi-color imaging. Finally, it is highly reproducible and scalable. While other methods have achieved some of these qualities, CUBIC is the first to realize all.
CUBIC provides information on previously unattainable 3D gene expression profiles and neural networks at the systems level. Because of its rapid and high-throughput imaging, CUBIC offers extraordinary opportunity to analyze localized effects of genomic editing. It also is expected to identify neural connections at the whole brain level. In fact, last author Hiroki Ueda is optimistic about further application to even larger mammalian systems. “In the near future, we would like to apply CUBIC technology to whole-body imaging at single cell resolution.”
Contact: Jens Wilkinson – RIKEN
Source: RIKEN press release
Image Source: The image is credited to Susaki et al/RIKEN/Cell and is adapted from the RIKEN press release
Original Research: Abstract for for “Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis” by Etsuo A. Susaki, Kazuki Tainaka, Dimitri Perrin, Fumiaki Kishino, Takehiro Tawara, Tomonobu M. Watanabe, Chihiro Yokoyama, Hirotaka Onoe, Megumi Eguchi, Shun Yamaguchi, Takaya Abe, Hiroshi Kiyonari, Yoshihiro Shimizu, Atsushi Miyawaki, Hideo Yokota and Hiroki R. Ueda. in Cell. Published online April 16 2014 doi:10.1016/j.cell.2014.03.042 Publication stage: In Press Corrected Proof
Important disclaimer: The abstract page offers this notice: “Note to users: Uncorrected proofs are Articles in Press that have been copy edited and formatted, but have not been finalized yet. They still need to be proof-read and corrected by the author(s) and the text could still change before final publication.






