Summary: A newly developed technique dubbed NeuroPAL is helping researchers investigate the dynamics of neural networks in the nervous system of microscopic worms.
Source: Columbia University
The human brain contains approximately 86 billion neurons, or nerve cells, woven together by an estimated 100 trillion connections, or synapses. Each cell has a role that helps us to move muscles, process our environment, form memories, and much more.
Given the huge number of neurons and connections, there is still much we don’t know about how neurons work together to give rise to thought or behavior.
Now Columbia scientists have engineered a coloring technique, known as NeuroPAL (a Neuronal Polychromatic Atlas of Landmarks), which makes it possible–at least in experiments with Caenorhabditis elegans (C. elegans), a worm species commonly used in biological research–to identify every single neuron in the mind of a worm.
Their research appears in the Jan. 7 issue of the journal Cell.
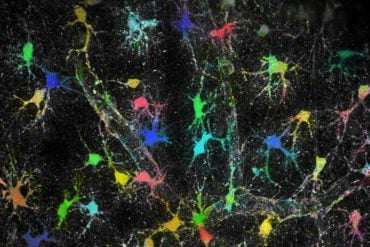
NeuroPAL, which uses genetic methods to “paint” neurons with fluorescent colors, permits, for the first time ever, scientists to identify each neuron in an animal’s nervous system, all while recording a whole nervous system in action.
“It’s amazing to ‘watch’ a nervous system in its entirety and see what it does,” said Oliver Hobert, professor in the Department of Biological Sciences at Columbia and a principal investigator with the Howard Hughes Medical Institute. “The images created are stunning– brilliant spots of color appear in the worm’s body like Christmas lights on a dark night.”
To conduct their research, the scientists created two software programs: one that identifies all the neurons in colorful NeuroPAL worm images and a second that takes the NeuroPAL method beyond the worm by designing optimal coloring for potential methods of identification of any cell type or tissue in any organism that permits genetic manipulations.
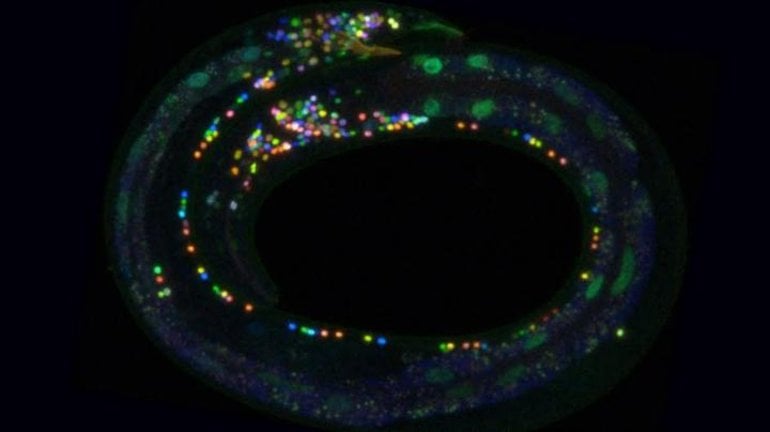
“We used NeuroPAL to record brainwide activity patterns in the worm and decode the nervous system at work,” said Eviatar Yemini, a postdoctoral researcher in the Department of Biological Sciences at Columbia and lead author of the study.
Because the colors are painted into the neuron’s DNA and linked to specific genes, the colors can also be used to reveal whether these specific genes are present or absent from a cell.
The researchers said that the novelty of the technique may soon be overshadowed by the discoveries it makes possible. In advance of their Cell publication, Hobert and Yemini released NeuroPAL to the scientific community, and several studies already have been published showing the utility of the tool.
“Being able to identify neurons, or other types of cells, using color can help scientists visually understand the role of each part of a biological system,” Yemini said. “That means when something goes wrong with the system, it may help pinpoint where the breakdown occurred.”
Collaborators on the study include Liam Paninski, Columbia University; Vivek Venkatachalam, Northeastern University; and Aravinthan Samuel, Harvard University.
About this brain mapping research news
Source: Columbia University
Contact: Carla Cantor – Columbia University
Image: The image is credited to Eviatar Yemini
Original Research: Closed access.
“NeuroPAL: A Multicolor Atlas for Whole-Brain Neuronal Identification in C. elegans” by Eviatar Yemini et al.Cell
Abstract
NeuroPAL: A Multicolor Atlas for Whole-Brain Neuronal Identification in C. elegans
Highlights
- •NeuroPAL: a strain with a stereotyped fluorescent color map to identify all neurons
- •NeuroPAL and semi-automated ID software pinpoint patterns of reporter gene expression
- •NeuroPAL identifies neuronal differentiation defects in mutant backgrounds
- •Dynamic whole-brain neuronal activity patterns defined by NeuroPAL in combination with GCaMP
Summary
Comprehensively resolving neuronal identities in whole-brain images is a major challenge. We achieve this in C. elegans by engineering a multicolor transgene called NeuroPAL (a neuronal polychromatic atlas of landmarks). NeuroPAL worms share a stereotypical multicolor fluorescence map for the entire hermaphrodite nervous system that resolves all neuronal identities. Neurons labeled with NeuroPAL do not exhibit fluorescence in the green, cyan, or yellow emission channels, allowing the transgene to be used with numerous reporters of gene expression or neuronal dynamics. We showcase three applications that leverage NeuroPAL for nervous-system-wide neuronal identification. First, we determine the brainwide expression patterns of all metabotropic receptors for acetylcholine, GABA, and glutamate, completing a map of this communication network. Second, we uncover changes in cell fate caused by transcription factor mutations. Third, we record brainwide activity in response to attractive and repulsive chemosensory cues, characterizing multimodal coding for these stimuli.