Summary: A new study uncovers the intricate molecular mechanisms behind schizophrenia by mapping genetic risk factors to specific brain cells. Researchers used comprehensive genetic and cellular analyses to identify how these genes affect different cell types in the prefrontal cortex.
The findings reveal that excitatory neurons are the most impacted, linking schizophrenia to neurodevelopment and synapse-related pathways. This research paves the way for targeted, personalized treatments for individuals with schizophrenia.
Key Facts:
- Cell Type Specificity: The study identifies which brain cell types express schizophrenia risk genes differently, highlighting excitatory neurons as the most affected.
- Pathway Implications: Transcriptional changes in these neurons implicate neurodevelopment and synaptic signaling pathways in schizophrenia.
- Personalized Treatments: Insights from the study could lead to tailored interventions, improving clinical outcomes for those with schizophrenia.
Source: McLean Hosptial
Schizophrenia is a complex disease with variable presentations, and the diverse nature of this mental health disorder has made understanding the mechanisms that cause the disease, and subsequently developing effective treatments, especially challenging.
In a new study, published May 23 is Science, a team led by McLean Hospital researchers used comprehensive genetic and cellular analyses to shed new light on the intricate molecular mechanisms underlying schizophrenia.
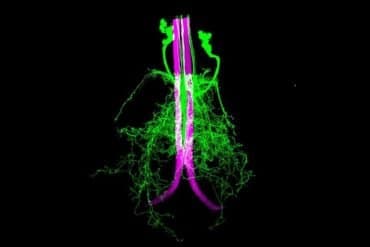
Their new work provides a map for how the genes known to increase risk of schizophrenia affect specific cells within the brain.
“We discovered which cell types express genes associated with schizophrenia risk differently, which biological functions are impacted within those cells, and which transcription factors are important for these changes,” explained lead and co-corresponding author, W. Brad Ruzicka MD, PhD, director of the Laboratory for Epigenomics in Human Psychopathology at McLean Hospital.
“This understanding will allow future treatments to be tailored to specific genes and cell types, as well as individuals with schizophrenia.”
Schizophrenia affects approximately 24 million people, or 1 in 300 people, worldwide, according to the World Health Organization.
For the new study, a multi-center team of researchers conducted a comprehensive single-cell analysis of transcriptomic changes in human prefrontal cortex, examining postmortem brain tissue from 140 individuals across two independent cohorts. Their analyses included more than 468,000 cells.
They uncovered unprecedented insights into the cellular basis of schizophrenia, linking genetic risk factors to specific neuronal populations. Specifically, the researchers found that excitatory neurons emerged as the most affected cell group, with transcriptional changes implicating neurodevelopment and synapse-related pathways.
Additionally, they found that known genetic risk factors for schizophrenia converge on alterations in specific neuronal populations, highlighting the interplay between rare and common genomic variants.
Through transcriptomic analysis, two distinct subpopulations of individuals with schizophrenia were identified, marked by the expression of specific excitatory and inhibitory neuronal cell states.
The new study suggests potential links between schizophrenia pathology and processes such as neurodevelopment, synaptic signaling, and transcriptional regulation, implicating key transcriptional regulators associated with both schizophrenia and neurodevelopmental disorders.
The study’s authors anticipate that insights gleaned from this research could pave the way for targeted interventions and personalized treatments for schizophrenia, potentially improving clinical outcomes for individuals affected by this debilitating and often disabling disorder.
The research team is now working to expand on these findings by investigating other regions of the brain and the molecular impact of other psychiatric diseases such as bipolar disorder.
They are also pursuing another dimension of complexity in this system by investigating isoform expression of implicated genes and how these cell type-specific gene expression changes lead to functional and potentially druggable changes in the protein space.
“This work advances understanding of schizophrenia pathophysiology at greater detail across both the complex landscape of cells within the brain, and the diverse experiences of people with this disease,” said Ruzicka, who is also associate medical director of Harvard Brain Tissue Resource Center at McLean, and an assistant professor of Psychiatry at Harvard Medical School.
“Our increased mechanistic understanding of schizophrenia provides avenues for future research to unravel the genetic and environmental underpinnings of this complex disease so we can provide our patients better care.”
Authorship: In addition to Ruzicka, co-authors from McLean include Sivan Subburaju; Daniel Reed Tso; and Makayla Hourihan.
Additional co-authors include Shahin Mohammadi; John F. Fullard; Jose Davila Velderrain; Shan Jiang6; Hao-Chih Lee; Jaroslav Bendl; PsychENCODE Consortium, Georgios Voloudakis; Vahram Haroutunian; Gabriel E. Hoffman; Panos Roussos; and Manolis Kellis.
Disclosures: The authors declare no competing interests.
Funding: This work was supported by National Institutes of Health (NIH) grant K08MH109759 to W.B.R.; Wilf Family Foundations to W.B.R. A full list of grants received by the authors can be found in the study.
About this genetics and schizophrenia research news
Author: Ryan Jaslow
Source: McLean Hospital
Contact: Ryan Jaslow – McLean Hospital
Image: The image is credited to Neuroscience News
Original Research: Closed access.
“Single-cell multi-cohort dissection of the schizophrenia transcriptome” by W. Brad Ruzicka et al. Science
Abstract
Single-cell multi-cohort dissection of the schizophrenia transcriptome
INTRODUCTION
Schizophrenia population genomics has identified strong germline genetic associations for this highly heritable disorder, and molecular investigation of postmortem brain samples has yielded evidence of transcriptomic and epigenomic alterations associated with this disease.
However, identifying molecular and cellular pathophysiological processes linking etiological risk factors and clinical presentation remains a challenge, due in part to the complex cellular architecture of the brain.
RATIONALE
Past work has implicated specific populations of excitatory and inhibitory neurons in the pathophysiology of schizophrenia, but existing large transcriptomic datasets of bulk tissue samples cannot directly assess cell type–specific contributions to disease.
Single-cell RNA sequencing technologies allow measurement of genome-wide gene expression in individual cells with high-throughput, moving beyond bulk tissue measures to map disease-associated transcriptional changes in discrete cellular populations without bias toward preselected cell types.
Investigating disease-associated phenotypic changes across the myriad cellular populations of the human brain can produce new insights into neuropsychiatric disease biology.
RESULTS
Using multiplexed single-nucleus RNA sequencing, we developed a single-cell resolution transcriptomic atlas of the prefrontal cortex across subjects with and without schizophrenia and present data from 468,727 nuclei isolated from 140 individuals across two well-defined and independently assayed cohorts.
We identified expression profiles of brain cell types and neuronal subpopulations and systematically characterized the transcriptional changes associated with schizophrenia in each.
For completeness, we report independent, cohort-specific analyses and joint meta-analysis of differential expression across 25 cell types. Using these data, we identified highly cell type–specific and reproducible expression changes, with 6634 differential expression events affecting 2455 genes and favoring down-regulated gene expression within excitatory neuronal populations.
We found significant overlap with previously reported bulk cortex expression changes, primarily for excitatory neuronal populations, whereas changes in lower-abundance cell types were less efficiently captured in tissue-level profiling. Differentially expressed genes enrich neurodevelopmental and synapse-related molecular pathways and point to a regulatory core of coexpressed transcription factors linked to genetic risk variants for schizophrenia and developmental delay.
Transcription factor targeting of schizophrenia differentially expressed genes in neuronal populations was validated with CUT&Tag in neuronal nuclei isolated from human prefrontal cortex.
Furthermore, both transcriptional changes and putative upstream regulatory factors were enriched with genes harboring common and rare risk variants for schizophrenia, presenting evidence that genetic risk variants across the population frequency spectrum tend to target genes with measurable expression alterations in the excitatory neurons of patients with schizophrenia.
Finally, the magnitude of schizophrenia-associated transcriptomic change segregated two populations of schizophrenia subjects. Transcriptomic heterogeneity within the cohorts was associated with specific cellular states shared across multiple neuronal populations, marked by genes related to synaptic function and one-carbon metabolism, suggesting genes characterizing distinct molecular phenotypes of schizophrenia.
CONCLUSION
Our results provide a valuable resource to investigate the molecular pathophysiology of schizophrenia at single-cell resolution, offering insights into preferential dysregulation of specific neuronal populations and their potential role in mediating genetic risk. Together, they suggest convergence of etiological genetic risk factors, neuronal transcriptional dysregulation, and symptomatic manifestation in schizophrenia.