Much like mapping the human genome laid the foundations for understanding the genetic basis of human health, new maps of the human epigenome may further unravel the complex links between DNA and disease. The epigenome is part of the machinery that helps direct how genes are turned off and on in different types of cells.
Researchers supported by the National Institutes of Health Common Fund’s Roadmap Epigenomics Program have mapped the epigenomes of more than 100 types of cells and tissues, providing new insight into which parts of the genome are used to make a particular type of cell. The data, available to the biomedical research community, can be found at the National Center for Biotechnology Information website.
“This represents a major advance in the ongoing effort to understand how the 3 billion letters of an individual’s DNA instruction book are able to instruct vastly different molecular activities, depending on the cellular context,” said NIH Director Francis Collins, M.D., Ph.D. “This outpouring of data-rich publications, produced by a remarkable team of creative scientists, provides powerful momentum for the rapidly growing field of epigenomics.”
Researchers from the NIH Common Fund’s Roadmap Epigenomics Program published a description of the epigenome maps in the journal Nature. More than 20 additional papers, published in Nature and Nature-associated journals, show how these maps can be used to study human biology.
“What the Roadmap Epigenomics Program has delivered is a way to look at the human genome in its living, breathing nature from cell type to cell type,” said Manolis Kellis, Ph.D., professor of computer science at the Massachusetts Institute of Technology, Cambridge, and senior author of the paper.
Understanding epigenomics
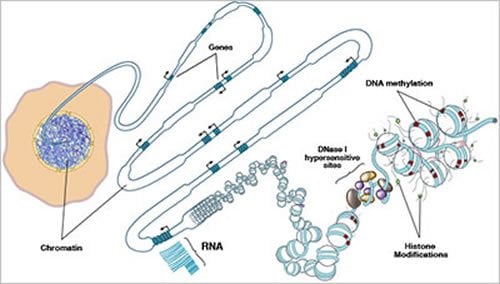
Almost all human cells have identical genomes that contain instructions on how to make the many different cells and tissues in the body. During the development of different types of cells, regulatory proteins turn genes on and off and, in doing so, establish a layer of chemical signatures that make up the epigenome of each cell. In the Roadmap Epigenomics Program, researchers compared these epigenomic signatures and established their differences across a variety of cell types. The resulting information can help us understand how changes to the genome and epigenome can lead to conditions such as Alzheimer’s disease, cancer, asthma, and fetal growth abnormalities.
The value of epigenomic data
Researchers can now take data from different cell types and directly compare them. “Today, sequencing the human genome can be done rapidly and cheaply, but interpreting the genome remains a challenge,” said Bing Ren, Ph.D., professor of cellular and molecular medicine at the University of California, San Diego, and co-author of the Nature paper and several of the associated papers. “These 111 reference epigenome maps are essentially a vocabulary book that helps us decipher each DNA segment in distinct cell and tissue types. These maps are like snapshots of the human genome in action.”

“This is the most comprehensive catalog of epigenomic data from primary human cells and tissues to date,” said Lisa Helbling Chadwick, Ph.D., project team leader and a program director at the National Institute of Environmental Health Sciences (NIEHS), part of NIH. “This coordinated effort, along with uniform data processing, makes it much easier for researchers to make direct comparisons across the entire data set.”
“Researchers from the 88 projects supported by the program, including those from this recent series of papers, have propelled the development of new epigenomic technologies,” said John Satterlee, Ph.D., co-coordinator of the Roadmap Epigenomics Program, and program director at the National Institute on Drug Abuse (NIDA), part of NIH. Satterlee added that the work of this program has served as a foundation for continued exploration of the human epigenome through the International Human Epigenome Consortium.
“With this increased understanding of the full epigenome, and the datasets available to the entire scientific community, the NIH Common Fund is striving to catalyze future research, to aid the understanding of how epigenomics plays a role in human diseases, with the expectation that further studies will identify early indications of disease and targets for therapeutics,” said James Anderson, M.D., Ph.D., director of NIH Division of Program Coordination, Planning, and Strategic Initiatives that oversees the NIH Common Fund.
NIDA, NIEHS, and the National Institute on Deafness and Other Communication Disorders are co-administrators of the NIH Common Fund’s Epigenomics Program.
Contact: Joe Balintfy – NIEHS/NIH
Source: NIEHS/NIH press release
Image Source: The top image is courtesy of Nature and Roadmap Epigenomics Consortium. The second image is courtesy of John Stamatoyannopoulos and Rae Senarighi. Both images are adapted from the NIEHS/NIH press release
Original Research: Full open access research for “Integrative analysis of 111 reference human epigenomes” by Roadmap Epigenomics Consortium, Anshul Kundaje, Wouter Meuleman, Jason Ernst, Misha Bilenky, Angela Yen, Alireza Heravi-Moussavi, Pouya Kheradpour, Zhizhuo Zhang, Jianrong Wang, Michael J. Ziller, Viren Amin, John W. Whitaker, Matthew D. Schultz, Lucas D. Ward, Abhishek Sarkar, Gerald Quon, Richard S. Sandstrom, Matthew L. Eaton, Yi-Chieh Wu, Andreas R. Pfenning, Xinchen Wang, Melina Claussnitzer, Yaping Liu, Cristian Coarfa, R. Alan Harris, Noam Shoresh, Charles B. Epstein, Elizabeta Gjoneska, Danny Leung, Wei Xie, R. David Hawkins, Ryan Lister, Chibo Hong, Philippe Gascard, Andrew J. Mungall, Richard Moore, Eric Chuah, Angela Tam, Theresa K. Canfield, R. Scott Hansen, Rajinder Kaul, Peter J. Sabo, Mukul S. Bansal, Annaick Carles, Jesse R. Dixon, Kai-How Farh, Soheil Feizi, Rosa Karlic, Ah-Ram Kim, Ashwinikumar Kulkarni, Daofeng Li, Rebecca Lowdon, GiNell Elliott, Tim R. Mercer, Shane J. Neph, Vitor Onuchic, Paz Polak, Nisha Rajagopal, Pradipta Ray, Richard C. Sallari, Kyle T. Siebenthall, Nicholas A. Sinnott-Armstrong, Michael Stevens, Robert E. Thurman, Jie Wu, Bo Zhang, Xin Zhou, Arthur E. Beaudet, Laurie A. Boyer, Philip L. De Jager, Peggy J. Farnham, Susan J. Fisher, David Haussler, Steven J. M. Jones, Wei Li, Marco A. Marra, Michael T. McManus, Shamil Sunyaev, James A. Thomson, Thea D. Tlsty, Li-Huei Tsai, Wei Wang, Robert A. Waterland, Michael Q. Zhang, Lisa H. Chadwick, Bradley E. Bernstein, Joseph F. Costello, Joseph R. Ecker, Martin Hirst, Alexander Meissner, Aleksandar Milosavljevic, Bing Ren, John A. Stamatoyannopoulos, Ting Wang and Manolis Kellis in Nature. Published online February 18 2015 doi:10.1038/nature14248