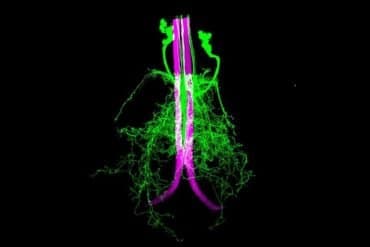
Summary: Dendrites weave their way through space that balances their need to connect to other neurons, but there is a cost.
Source: University of Oregon
High-resolution imaging and 3D computer modeling show that the dendrites of neurons weave through space in a way that balances their need to connect to other neurons with the costs of doing so.
The discovery, reported in Nature Scientific Reports Jan. 27, emerged as researchers sought to understand the fractal nature of neurons as part of a University of Oregon project to design fractal-shaped electrodes to connect with retinal neurons to address vision loss due to retinal diseases.
“The challenge in our research has been understanding how the neurons we want to target in the retina will connect to our electrodes,” said Richard Taylor, a professor and head of the UO physics department. “Essentially, we have to fool the neurons into thinking that the electrode is another neuron by making the two have the same fractal character.”
Working with collaborators at the University of Auckland and University of Canterbury in New Zealand, confocal microscopy of neurons in the hippocampal region of a rat’s brain revealed an intricate interplay of branches weaving through space at multiple size scales before connecting to other neurons. That, Taylor said, raised the question, why adopt such a complicated pattern?
With the help of UO post-doctoral researcher Saba Moslehi, doctoral students Julian H. Smith and Conor Rowland turned to 3D modeling to explore what happens when they manipulated the dendrites of more than 1,600 neurons into unnatural forms, straightening them or curling them up.
“By distorting their branches and looking at what happens, we were able to show that the fractal weaving of the natural branches is balancing the ability of neurons to connect with their neighbors to form natural electric circuits while balancing the construction and operating costs of the circuits,” Rowland said.
Using a fractal analysis known as the box-counting technique, the researchers were able to assign fractal dimensions, or D values, that quantify the relative contributions of the coarse- and fine-scaled dendrites to a neuron’s fractal pattern. These D values, Taylor said, will be important in optimizing his team’s tiny electrodes for implanting at the back of eyes to stimulate retinal neurons.
“Our implants will have to accommodate the neurons’ weaving branches through careful selection of their D values,” said Taylor, a member of the UO’s Materials Science Institute. “Unlike building a straight runway so a pilot can land efficiently, our electrodes will need to act like a weaving runway so that the neurons can connect without changing their behavior.”

Nature’s fractals benefit from how they grow at multiple scales, said Taylor, who has long turned to fractals as bioinspiration. While trees have the most-recognized form of fractal branching, this work, he said, highlights how neurons are different from trees.
“Whereas the fractal character of trees originates predominantly from the distribution of branch sizes, the neurons also use the way their branches weave through space to generate their fractal character,” Taylor said.
Taylor, a Cottrell Scholar of the Research Council for Science Advancement, was granted a sweeping U.S. patent in 2015 for not only his development of artificial fractal-based implants related to vision but also to all such implants that link signaling activity with nerves for any purpose in animal and human biology.
Taylor and co-authors closed their paper by raising the possibility that the D values of neuronal networking may benefit research on numerous brain-related diseases. For Alzheimer’s disease, Taylor said, D values could be a measure for understanding declines in connectivity between neurons.
“A lot of diseases result in losing connectivity, and neuron D values may be dropping as they move into a pathological state,” he said.
Additional UO co-authors on the paper were doctoral students Rick D. Montgomery, Kris Schobert and William Watterson. Other co-authors were Bruce Harland, a research fellow at the University of Auckland, and John Dalrymple-Alford, a neuroscientist at the University of Canterbury.
Funding: The W.M. Keck Foundation, Living Legacy Foundation and Ciminelli Foundation funded the research.
About this neuroscience research news
Source: University of Oregon
Contact: Jim Barlow – University of Oregon
Image: The image is credited to Richard Taylor
Original Research: Open access.
“How neurons exploit fractal geometry to optimize their network connectivity” by Julian H. Smith, Conor Rowland, B. Harland, S. Moslehi, R. D. Montgomery, K. Schobert, W. J. Watterson, J. Dalrymple-Alford & R. P. Taylor. Scientific Reports
Abstract
How neurons exploit fractal geometry to optimize their network connectivity
We investigate the degree to which neurons are fractal, the origin of this fractality, and its impact on functionality. By analyzing three-dimensional images of rat neurons, we show the way their dendrites fork and weave through space is unexpectedly important for generating fractal-like behavior well-described by an ‘effective’ fractal dimension D.
This discovery motivated us to create distorted neuron models by modifying the dendritic patterns, so generating neurons across wide ranges of D extending beyond their natural values. By charting the D-dependent variations in inter-neuron connectivity along with the associated costs, we propose that their D values reflect a network cooperation that optimizes these constraints. We discuss the implications for healthy and pathological neurons, and for connecting neurons to medical implants.
Our automated approach also facilitates insights relating form and function, applicable to individual neurons and their networks, providing a crucial tool for addressing massive data collection projects (e.g. connectomes).