Summary: New findings about NLGN4 genes reveals why males are more susceptible to developing autism than females.
Source: NIH-NINDS
A new study in Neuron offers clues to why autism spectrum disorder (ASD) is more common in boys than in girls. National Institutes of Health scientists found that a single amino acid change in the NLGN4 gene, which has been linked to autism symptoms, may drive this difference in some cases. The study was conducted at NIH’s National Institute of Neurological Disorders and Stroke (NINDS).
Researchers led by Katherine Roche, Ph.D., a neuroscientist at NINDS, compared two NLGN4 genes, (one on the X chromosome and one on the Y chromosome), which are important for establishing and maintaining synapses, the communication points between neurons.
Every cell in our body contains two sex chromosomes. Females have two X chromosomes; males have one X and one Y chromosome. Until now, it was assumed that the NLGN4X and NLGN4Y genes, which encode proteins that are 97% identical, functioned equally well in neurons.
But using a variety of advanced technology including biochemistry, molecular biology, and imaging tools, Dr. Roche and her colleagues discovered that the proteins encoded by these genes display different functions. The NLGN4Y protein is less able to move to the cell surface in brain cells and is therefore unable to assemble and maintain synapses, making it difficult for neurons to send signals to one another. When the researchers fixed the error in cells in a dish, they restored much of its correct function.
“We really need to look at NLGN4X and NLGN4Y more carefully,” said Thien A. Nguyen, Ph.D., first author of the study and former graduate student in Dr. Roche’s lab. “Mutations in NLGN4X can lead to widespread and potentially very severe effects in brain function, and the role of NLGNY is still unclear.”
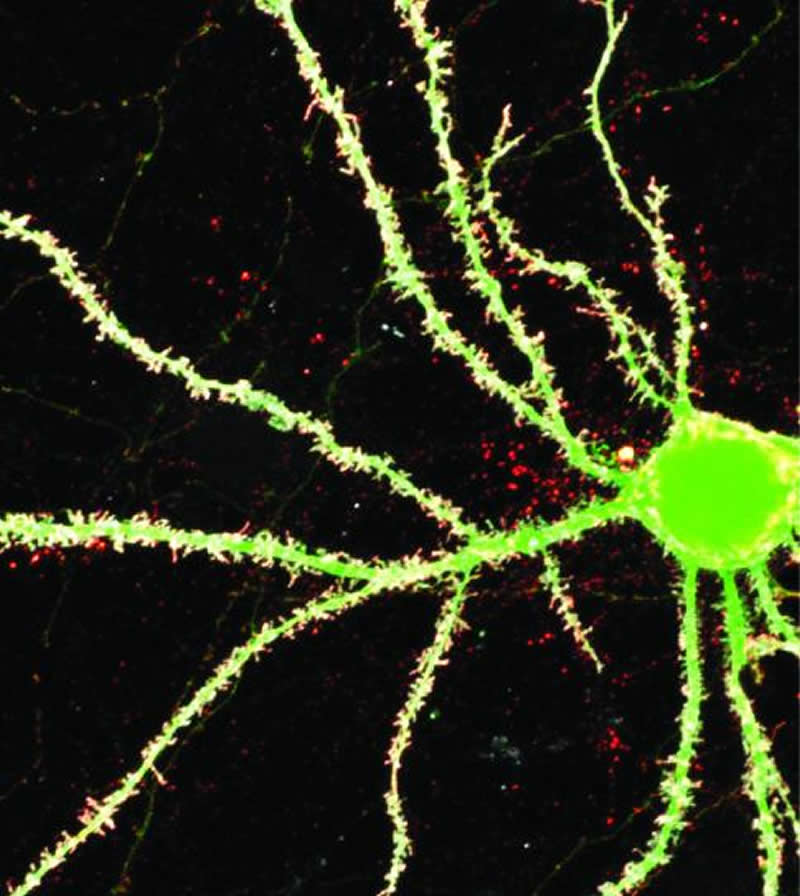
Dr. Roche’s team found that the problems with NLGN4Y were due to a single amino acid. The researchers also discovered that the region surrounding that amino acid in NLGN4X is sensitive to mutations in the human population. There are a cluster of variants found in this region in people with ASD and intellectual disability and these mutations result in a deficit in function for NLGN4X that is indistinguishable from NLGN4Y.
In females, when one of the NLGN4X genes has a mutation, the other one can often compensate. However, in males, diseases can occur when there is a mutation in NLGN4X because there is no compensation from NLGN4Y.
The current study suggests that if there is a mutation in NLGN4X, NLGN4Y is not able to take over, because it is a functionally different protein. If the mutations occur in regions of NLGN4X that affect the protein levels, that may result in autism-related symptoms including intellectual deficits. The inability of NLGN4Y to compensate for mutations in NLGN4X may help explain why males, who only have one X chromosome, tend to have a greater incidence of NLGN4X-associated ASD than females.
“The knowledge about these proteins will help doctors treating patients with mutations in NLGN4X better understand their symptoms,” said Dr. Roche.
Funding: This work was supported by the NIH Intramural Research Program.
Source:
NIH-NINDS
Media Contacts:
Barbara McMakin – NIH-NINDS
Image Source:
The image is credited to Roche Lab/NINDS.
Original Research: Closed access
“A Cluster of Autism-Associated Variants on X-Linked NLGN4X Functionally Resemble NLGN4Y”. Thien A. Nguyen, Kunwei Wu, Saurabh Pandey, Alexander W. Lehr, Yan Li, Michael A. Bemben, John D. Badger II, Julie L. Lauzon, and others.
Neuron doi:10.1016/j.neuron.2020.03.008.
Abstract
A Cluster of Autism-Associated Variants on X-Linked NLGN4X Functionally Resemble NLGN4Y
Highlights
• Despite sharing ∼97% amino acid identity, NLGN4X and NLGN4Y are differentially regulated
• NLGN4Y cannot traffic to the surface due to one amino difference from NLGN4X
• A cluster of autism-associated variants in NLGN4X surrounds the critical amino acid
• NLGN4X autism-associated variants display a deficit in trafficking similar to NLGN4Y
Summary
Autism spectrum disorder (ASD) is more prevalent in males; however, the etiology for this sex bias is not well understood. Many mutations on X-linked cell adhesion molecule NLGN4X result in ASD or intellectual disability. NLGN4X is part of an X-Y pair, with NLGN4Y sharing ∼97% sequence homology. Using biochemistry, electrophysiology, and imaging, we show that NLGN4Y displays severe deficits in maturation, surface expression, and synaptogenesis regulated by one amino acid difference with NLGN4X. Furthermore, we identify a cluster of ASD-associated mutations surrounding the critical amino acid in NLGN4X, and these mutations phenocopy NLGN4Y. We show that NLGN4Y cannot compensate for the functional deficits observed in ASD-associated NLGN4X mutations. Altogether, our data reveal a potential pathogenic mechanism for male bias in NLGN4X-associated ASD.