Summary: Mouse pups with the autism-associated 16p11.2 deletion can not able to correctly perform a specific vocalization pattern to call for their mother when separated from her in the same way those without the deletion can, a new study reveals.
Source: UT Health San Antonio
One-fifth of babies who inherit a genetic variant located on chromosome 16 will develop autism spectrum disorder (ASD) by age 3. The variant is called 16p11.2 deletion.
Noboru Hiroi, PhD, of The University of Texas Health Science Center at San Antonio (also referred to as UT Health San Antonio), is studying mice that have this deletion. The team, which includes colleagues from Japan, Ireland and the U.S., is harnessing the power of machine learning to understand which vocalizations of the newborn mouse pups are most predictive of social abnormalities one month later when the pups reach puberty.
“It is essential to identify those very early signs that can predict what is to come, because if we can translate what we discover in mouse pups to human infants and apply therapeutic options earlier, their outcome will be better,” Dr. Hiroi said.
Mice are studied because of the short timeline of their development. The same research in human babies, which is just beginning, will take two to three years of follow-up. In May, collaborators led by Cynthia Blanco, MD, professor of pediatrics and the Greehey Family Foundation Chair in Neonatology Research at UT Health San Antonio, started recording the cries of preterm infants in the University Hospital neonatal intensive care unit. Preterm infants are targeted because of their high risk of developmental issues, including ASD.
Disordered cries
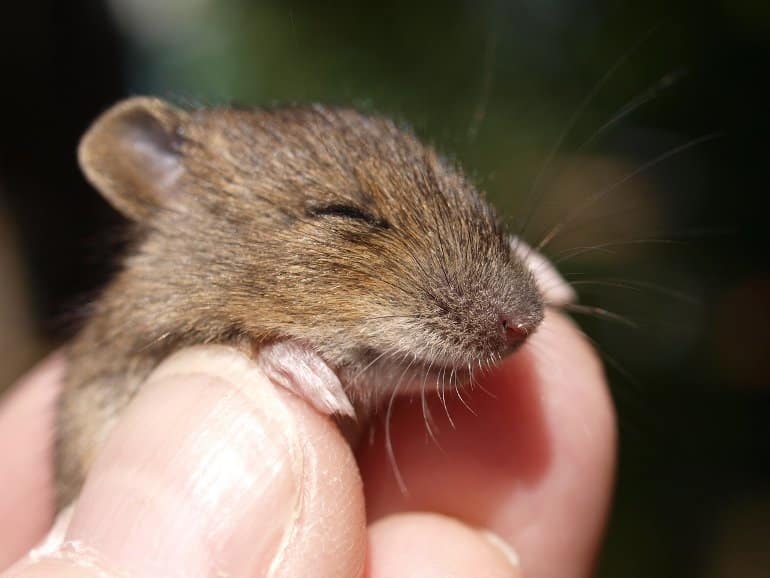
When mouse pups are separated from their mothers, they emit ultrasonic vocalizations in a certain order. Mouse mothers respond to the cries and take care of their offspring.
The cries are abnormal in mice with the 16p11.2 deletion.
“Pups that carry this genetic variation cannot form the correct sequence,” Dr. Hiroi said. “We want to know whether those abnormal sequences or combinations of call types can predict what is to come one month later in their social behaviors.”
Machine learning will enable the team to develop a precise diagnostic algorithm for ASD. “The point is, once we can do this with the mouse vocalizations, we can apply the same algorithm to the cries of human babies,” Dr. Hiroi said.
Earlier intervention
Infants at risk of ASD who are identified in this way can be given desensitization therapy so that they don’t overreact to certain cues they don’t like, he said. And behavioral therapies can be applied to help babies learn how to cope in social situations.

The research is described in a paper published in the journal Molecular Psychiatry.
The authors point out that algorithms developed from infant vocalizations could be used to predict other disorders, such as schizophrenia, which develops 17 to 20 years after birth.
The mice also display abnormality in their responses to social olfactory cues. Normal mice presented with another mouse respond by sniffing. The mutant mice don’t respond to that cue, Dr. Hiroi said.
Acknowledgments
Dr. Hiroi, recruited to UT Health San Antonio in 2019 from the Albert Einstein College of Medicine, is a professor in the departments of pharmacology, cellular and integrative physiology, and cell systems and anatomy within the university’s Joe R. and Teresa Lozano Long School of Medicine.
Funding: This research is made possible by a grant (R01DC015776) to Dr. Hiroi from the National Institute on Deafness and Other Communication Disorders, part of the National Institutes of Health. The Department of Psychiatry and Behavioral Sciences at UT Health San Antonio also provided startup funds for this project.
About this autism research news
Source: UT Health San Antonio
Contact: Will Sansom – UT Health San Antonio
Image: The image is in the public domain
Original Research: Open access.
“Computational identification of variables in neonatal vocalizations predictive for postpubertal social behaviors in a mouse model of 16p11.2 deletion” by Mitsuteru Nakamura, Kenny Ye, Mariel Barbachan e Silva, Takahira Yamauchi, Daniel J. Hoeppner, Amir Fayyazuddin, Gina Kang, Emi A. Yuda, Masako Nagashima, Shingo Enomoto, Takeshi Hiramoto, Richard Sharp, Itaru Kaneko, Katsunori Tajinda, Megumi Adachi, Takuma Mihara, Shinichi Tokuno, Mark A. Geyer, Pilib Ó Broin, Mitsuyuki Matsumoto and Noboru Hiroi. Molecular Psychiatry
Abstract
Computational identification of variables in neonatal vocalizations predictive for postpubertal social behaviors in a mouse model of 16p11.2 deletion
Autism spectrum disorder (ASD) is often signaled by atypical cries during infancy. Copy number variants (CNVs) provide genetically identifiable cases of ASD, but how early atypical cries predict a later onset of ASD among CNV carriers is not understood in humans. Genetic mouse models of CNVs have provided a reliable tool to experimentally isolate the impact of CNVs and identify early predictors for later abnormalities in behaviors relevant to ASD.
However, many technical issues have confounded the phenotypic characterization of such mouse models, including systematically biased genetic backgrounds and weak or absent behavioral phenotypes.
To address these issues, we developed a coisogenic mouse model of human proximal 16p11.2 hemizygous deletion and applied computational approaches to identify hidden variables within neonatal vocalizations that have predictive power for postpubertal dimensions relevant to ASD. After variables of neonatal vocalizations were selected by least absolute shrinkage and selection operator (Lasso), random forest, and Markov model, regression models were constructed to predict postpubertal dimensions relevant to ASD.
While the average scores of many standard behavioral assays designed to model dimensions did not differentiate a model of 16p11.2 hemizygous deletion and wild-type littermates, specific call types and call sequences of neonatal vocalizations predicted individual variability of postpubertal reciprocal social interaction and olfactory responses to a social cue in a genotype-specific manner.
Deep-phenotyping and computational analyses identified hidden variables within neonatal social communication that are predictive of postpubertal behaviors.