Summary: Research reveals atrophy of the amygdala and hippocampus at age 40 in patients later diagnosed with Alzheimer’s disease.
Source: CNRS – French National Centre for Scientific Research
What changes in the brain are caused by Alzheimer’s disease? How do these changes differ from those observed in the normal ageing process? Researchers from the CNRS, the École pratique des hautes études (EPHE) and the University of Valencia (Spain) explored these questions by analysing over 4,000 MRI scans of healthy and diseased brains using the “volBrain” platform. Their models, published in the March 8, 2019 edition of Scientific Reports, reveal an early atrophy of the amygdala and hippocampus at age 40 in patients with Alzheimer’s disease.
Alzheimer’s disease is characterised by changes in the brain, some of which can be measured in the patient using biomarkers such as the size of different areas of the brain. Studying these biomarkers has shown that certain brain structures shrink in the presence of Alzheimer’s well before the first signs of dementia appear. Key questions remained however: when and how did changes in these biomarkers differ between a healthy subject and one with Alzheimer’s, for example?
To answer these questions, researchers at the Laboratoire bordelais de recherche en informatique (LaBRI) (CNRS/Bordeaux INP/Université de Bordeaux), at the Aquitaine Institute for Cognitive and Integrative Neuroscience (CNRS/Université de Bordeaux/EPHE) and the University of Valencia (Spain) created a model which traces brain changes and activity over an entire life span using a massive set of over 4,000 MRI scans processed with the volBrain[1] platform. Prior to this, scientists did not have images covering every period of an Alzheimer patient’s life. Researchers suggested modelling the changes generally seen in the volumes of different structures using a vast quantity of samples in order to pinpoint where healthy brains diverged from diseased ones over time.
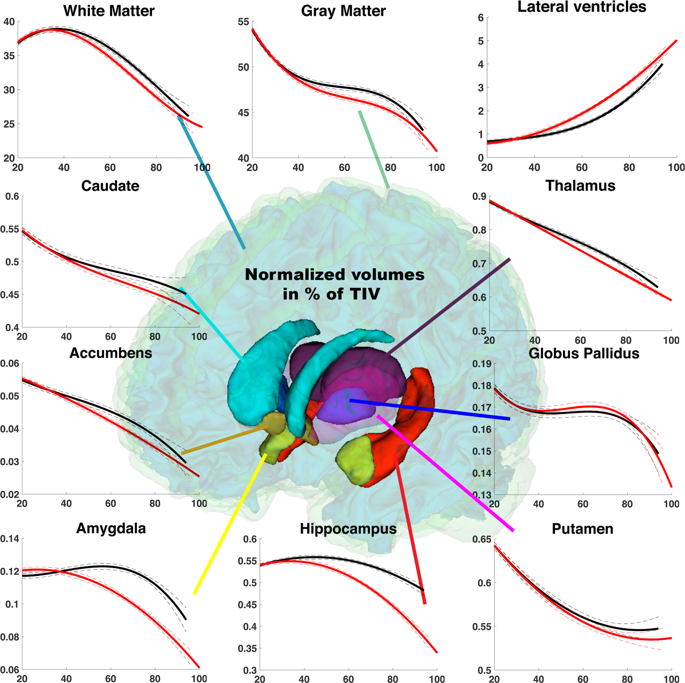
Based on MRI scans from 2,944 healthy control subjects between the ages of 9 months to 94 years old, the team developed a ‘normal’ model of average brain changes, which they compared to a pathological model based on MRIs from 1,385 Alzheimer’s patients aged over 55 and 1,877 young control subjects. Their results show an early divergence between the pathological models and the normal trajectory of ageing of the hippocampus before age 40, and of the amygdala around age 40. Both of these structures suffer atrophy in the presence of Alzheimer’s disease. Also evident is an early enlargement, in patients with the disease, of an internal cavity in the brain known as the lateral ventricle. This enlargement is part of the ageing process in normal subjects too, however, thus limiting the pertinence of this measurement in subjects of an advanced age, and reaffirming the usefulness of studying biomarkers across an entire life span.
Developed by the CNRS, the Université de Bordeaux and the University of Valencia in Spain, this free platform allows researchers around the world to upload structural MRI files and obtain in record time an automatic analysis of the scanned brain structure volumes. Since its creation in 2015, over 110,000 brain MRIs have been analysed for over 2,500 users worldwide.
Source:
CNRS
Media Contacts:
Alexiane Agullo – CNRS
Image Source:
Image from research paper listed below.
Original Research: Open Access
“Lifespan Changes of the Human Brain In Alzheimer’s Disease” by Pierrick Coupé, José Vicente Manjón, Enrique Lanuza & Gwenaelle Catheline in Scientific Reports volume 9, Article number: 3998 (2019)
DOI: 10.1038/s41598-019-39809-8
Abstract
Lifespan Changes of the Human Brain In Alzheimer’s Disease
Brain imaging studies have shown that slow and progressive cerebral atrophy characterized the development of Alzheimer’s Disease (AD). Despite a large number of studies dedicated to AD, key questions about the lifespan evolution of AD biomarkers remain open. When does the AD model diverge from the normal aging model? What is the lifespan trajectory of imaging biomarkers for AD? How do the trajectories of biomarkers in AD differ from normal aging? To answer these questions, we proposed an innovative way by inferring brain structure model across the entire lifespan using a massive number of MRI (N = 4329). We compared the normal model based on 2944 control subjects with the pathological model based on 3262 patients (AD + Mild cognitive Impaired subjects) older than 55 years and controls younger than 55 years. Our study provides evidences of early divergence of the AD models from the normal aging trajectory before 40 years for the hippocampus, followed by the lateral ventricles and the amygdala around 40 years. Moreover, our lifespan model reveals the evolution of these biomarkers and suggests close abnormality evolution for the hippocampus and the amygdala, whereas trajectory of ventricular enlargement appears to follow an inverted U-shape. Finally, our models indicate that medial temporal lobe atrophy and ventricular enlargement are two mid-life physiopathological events characterizing AD brain.