A Dartmouth researcher and his colleagues have discovered a way to predict human emotions based on brain activity.
The study is unusual because of its accuracy — more than 90 percent — and the large number of participants who reflect the general adult population rather than just college students. The findings could help in diagnosing and treating a range of mental and physical health conditions.
The study appears in the journal PLOS Biology.
“It’s an impressive demonstration of imaging our feelings, of decoding our emotions from brain activity,” says lead author Luke Chang, an assistant professor in Psychological and Brain Sciences at Dartmouth. “Emotions are central to our daily lives and emotional dysregulation is at the heart of many brain- and body-related disorders, but we don’t have a clear understanding of how emotions are processed in the brain. Thus, understanding the neurobiological mechanisms that generate and reduce negative emotional experiences is paramount.”
The quest to understand the “emotional brain” has motivated hundreds of neuroimaging studies in recent years. But for neuroimaging to be useful, sensitive and specific “brain signatures” must be developed that can be applied to individual people to yield information about their emotional experiences, neuropathology or treatment prognosis. Thus far, the neuroscience of emotion has yielded many important results but no such indicators for emotional experiences.
In their new study, the researchers’ goals were to develop a brain signature that predicts the intensity of negative emotional responses to evocative images; to test the signature in generalizing across individual participants and images; to examine the signature’s specificity related to pain; and to explore the neural circuitry necessary to predict negative emotional experience.
Chang and his colleagues studied 182 participants who were shown negative photos (bodily injuries, acts of aggression, hate groups, car wrecks, human feces) and neutral photos. Thirty additional participants were also subjected to painful heat. Using brain imaging and machine learning techniques, the researchers identified a neural signature of negative emotion — a single neural activation pattern distributed across the entire brain that accurately predicts how negative a person will feel after viewing unpleasant images.
“This means that brain imaging has the potential to accurately uncover how someone is feeling without knowing anything about them other than their brain activity,” Chang says. “This has enormous implications for improving our understanding of how emotions are generated and regulated, which have been notoriously difficult to define and measure. In addition, these new types of neural measures may prove to be important in identifying when people are having abnormal emotional responses – for example, too much or too little — which might indicate broader issues with health and mental functioning.”
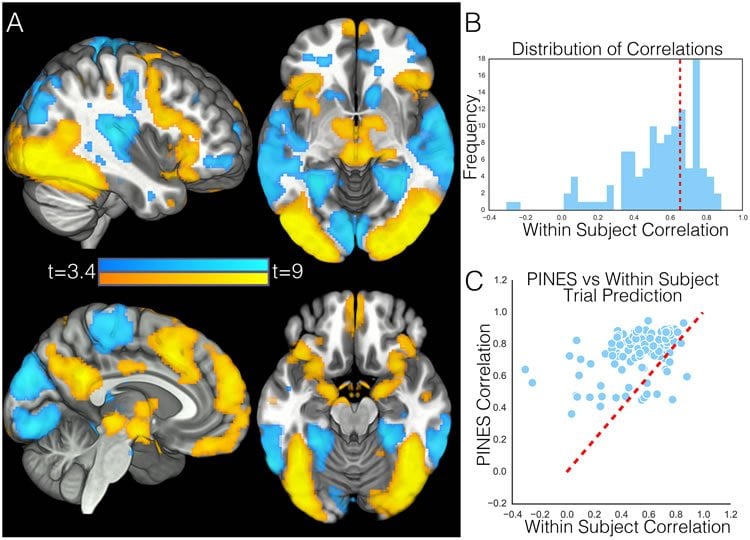
Unlike most previous research, the new study included a large sample size that reflects the general adult population and not just young college students; used machine learning and statistics to develop a predictive model of emotion; and, most importantly, tested participants across multiple psychological states, which allowed researchers to assess the sensitivity and specificity of their brain model.
“We were particularly surprised by how well our pattern performed in predicting the magnitude and type of aversive experience,” Chang says. “As skepticism for neuroimaging grows based on over-sold and -interpreted findings and failures to replicate based on small sizes, many neuroscientists might be surprised by how well our signature performed. Another surprising finding is that our emotion brain signature using lots of people performed better at predicting how a person was feeling than their own brain data. There is an intuition that feelings are very idiosyncratic and vary across people. However, because we trained the pattern using so many participants – for example, four to 10 times the standard fMRI experiment — we were able to uncover responses that generalized beyond the training sample to new participants remarkably well.”
Funding: The study included researchers from the University of Colorado and University of Pittsburgh and was supported by the National Institute of Health.
Source: John Cramer – Dartmouth College
Image Source: The image is credited to Chang et al./PLOS Biology
Original Research: Full open access research for “A Sensitive and Specific Neural Signature for Picture-Induced Negative Affect” by Luke J. Chang, Peter J. Gianaros, Stephen B. Manuck, Anjali Krishnan, and Tor D. Wager in PLOS Biology. Published online June 22 2015 doi:10.1371/journal.pbio.1002180
Abstract
A Sensitive and Specific Neural Signature for Picture-Induced Negative Affect
Neuroimaging has identified many correlates of emotion but has not yet yielded brain representations predictive of the intensity of emotional experiences in individuals. We used machine learning to identify a sensitive and specific signature of emotional responses to aversive images. This signature predicted the intensity of negative emotion in individual participants in cross validation (n =121) and test (n = 61) samples (high–low emotion = 93.5% accuracy). It was unresponsive to physical pain (emotion–pain = 92% discriminative accuracy), demonstrating that it is not a representation of generalized arousal or salience. The signature was comprised of mesoscale patterns spanning multiple cortical and subcortical systems, with no single system necessary or sufficient for predicting experience. Furthermore, it was not reducible to activity in traditional “emotion-related” regions (e.g., amygdala, insula) or resting-state networks (e.g., “salience,” “default mode”). Overall, this work identifies differentiable neural components of negative emotion and pain, providing a basis for new, brain-based taxonomies of affective processes.
“A Sensitive and Specific Neural Signature for Picture-Induced Negative Affect” by Luke J. Chang, Peter J. Gianaros, Stephen B. Manuck, Anjali Krishnan, and Tor D. Wager in PLOS Biology. Published online June 22 2015 doi:10.1371/journal.pbio.1002180






