Summary: Researchers identify the same mutation in different areas of the genome that cause at least three different diseases.
Source: University of Tokyo
Families living with extremely rare neurodegenerative diseases finally received an answer to the cause of their illnesses, thanks to a researcher’s hunch and decades of improvements in DNA sequencing technology.
Four different rare diseases are all caused by the same short segment of DNA repeated too many times, a mutation researchers call noncoding expanded tandem repeats. Researchers suspect variations of this type of mutation may cause other diseases that have thus far evaded diagnosis by genetic testing.
Treatment strategies for many genetic diseases are complicated by the fact that different mutations can cause the same disease. For example, Parkinson’s disease can be caused by unique mutations in at least five different genes. Cystic fibrosis can be caused by over 1,000 different mutations in the same gene.
Researchers are excited because instead of finding unique mutations in specific genes, they identified the same mutation in different areas of the genome causing different diseases.
Genetic cause of rare diseases evaded diagnosis
“Because the mutations causing the diseases are so similar, in the future, all these patients might benefit from the same treatment,” explained Dr. Hiroyuki Ishiura, M.D./Ph.D., assistant professor from the University of Tokyo Hospital and first author of the recent research paper published in Nature Genetics.
“Gene silencing techniques [which inactivate previously active genes] are a possible treatment. We cannot know the result, but we believe such strategies may help patients in the future,” said Dr. Shoji Tsuji, M.D./Ph.D., project professor from the University of Tokyo Hospital, a corresponding author of the recent research paper.
The research team focused on patients with adult-onset neurodegeneration, showing symptoms like cognitive impairment, uncontrolled movement, loss of balance, weakness in the arms and legs, or difficulty swallowing.
The genetic cause of one disease with those symptoms, fragile X tremor/ataxia syndrome, was identified in the early 2000s as three letters of the genetic code, CGG, being repeated dozens or hundreds of times on the X chromosome. Noncoding expanded tandem repeat mutations can be caused by any letters of the genetic code repeated an unusual number of times anywhere in the genome.
Researchers had a hunch that the same CGG repeat mutation might cause three other rare diseases with similar symptoms and clinical test results. But since patients with those other diseases had normal X chromosomes, researchers had no idea where in the genome the potential CGG repeat mutations might exist.
High-tech scavenger hunt
Previous generations of DNA sequencing technology required researchers to know where in the genome to look for a mutation. Searching for CGG repeat mutations on all 46 chromosomes in the human genome was extremely difficult and laborious.
The new approach that researchers designed relies on modern next-generation genome sequencing and clever data analysis.
Researchers sequenced patients’ and healthy people’s entire genomes in short, overlapping but broken stretches. Collaborators, including Professor Shinichi Morishita, a computational biologist from the Department of Bioinformatics and Systems Biology, developed a new computer program to sort all of those short sequences and search for ones made of just CGG over and over again.
Using a standard sequence of the entire healthy human genome, the computer program could pin down those short segments containing CGG repeat mutations to particular genetic neighborhoods. Researchers narrowed down their search to anywhere in the genome where patients had a large number of CGG repeats and healthy people had none.
With that information, researchers then knew where to sequence a long stretch of DNA to specifically identify the gene and wherein the gene the patients’ CGG repeat mutations exist.
Rare diseases share common symptoms
All four rare neurological diseases that the research team studied are caused by CGG repeat mutations in distant, seemingly unrelated areas of the genome.
One of the diseases is currently only known in a single-family.
“We cared for this family starting 10 years ago, so it was a long puzzle for us to find the correct diagnosis,” said Ishiura.
Efforts are ongoing to study genomes identified through the genetic analysis of people who are healthy despite having the same CGG repeat mutations. Such cases are extremely rare.
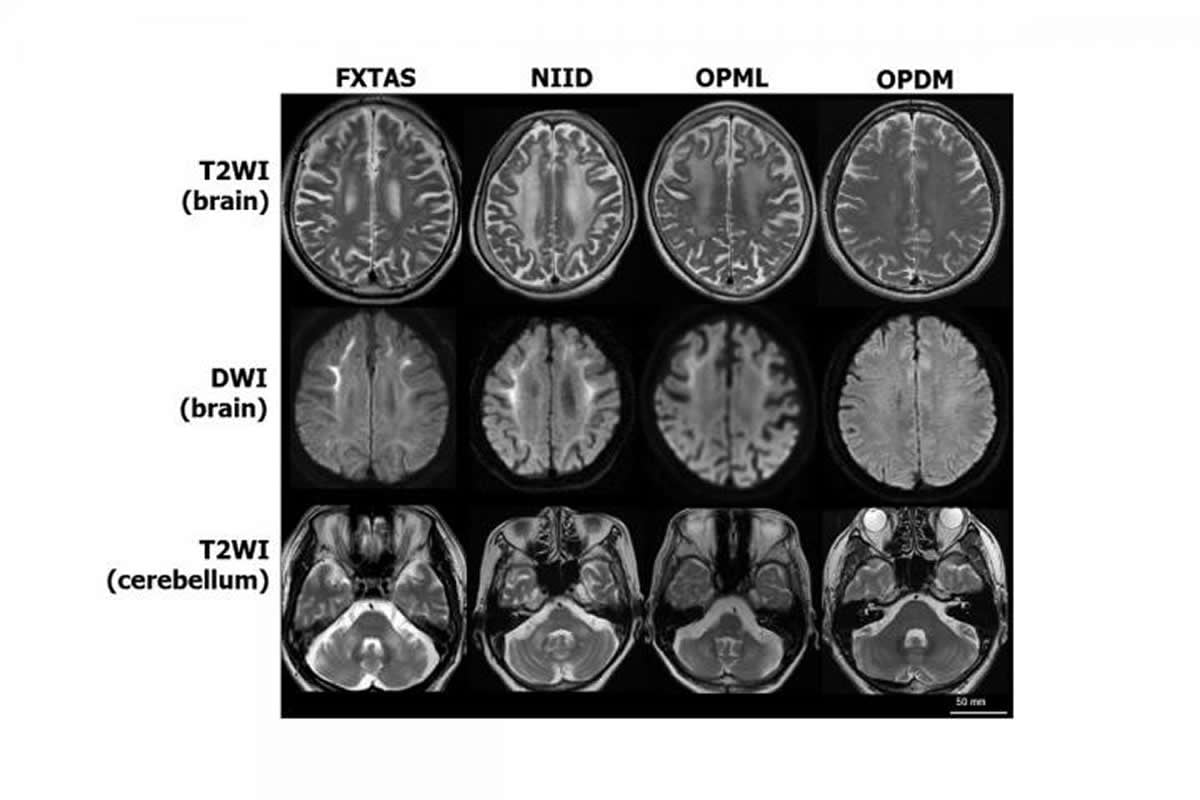
The researchers hope that their studies on rare neurodegenerative diseases might lead to insights into more common diseases caused by other types of noncoding tandem repeat mutations, including amyotrophic lateral sclerosis (ALS), also known as motor neuron disease or Lou Gehrig’s disease.
In 2011, Tsuji received the Japanese government’s Medal with Purple Ribbon for contributions to academic development in the field of intractable neurological diseases. In 2015, he was awarded the medal for scientific achievement by the World Federation of Neurology for his leadership in the field of neurogenetics.
Source:
University of Tokyo
Media Contacts:
Shoji Tsuji, M.D./Ph.D – University of Tokyo
Image Source:
The image is credited to Hiroyuki Ishiura and Shoji Tsuji.
Original Research: Closed access
“Noncoding CGG repeat expansions in neuronal intranuclear inclusion disease, oculopharyngodistal myopathy and an overlapping disease”. Hiroyuki Ishiura, Shota Shibata, Jun Yoshimura, Yuta Suzuki, Wei Qu, Koichiro Doi, M. Asem Almansour, Junko Kikuchi, Makiko Taira, Jun Mitsui, Yuji Takahashi, Yaeko Ichikawa, Tatsuo Mano, Atsushi Iwata, Yasuo Harigaya, Miho Kawabe Matsukawa, Takashi Matsukawa, Masaki Tanaka, Yuichiro Shirota, Ryo Ohtomo, Hisatomo Kowa, Hidetoshi Date, Aki Mitsue, Hiroyuki Hatsuta, Satoru Morimoto, Shigeo Murayama, Yasushi Shiio, Yuko Saito, Akihiko Mitsutake, Mizuho Kawai, Takuya Sasaki, Yusuke Sugiyama, Masashi Hamada, Gaku Ohtomo, Yasuo Terao, Yoshihiko Nakazato, Akitoshi Takeda, Yoshio Sakiyama, Yumi Umeda-Kameyama, Jun Shinmi, Katsuhisa Ogata, Yutaka Kohno, Shen-Yang Lim, Ai Huey Tan, Jun Shimizu, Jun Goto, Ichizo Nishino, Tatsushi Toda, Shinichi Morishita, Shoji Tsuji
Nature Genetics. doi:10.1038/s41588-019-0458-z
Abstract
Noncoding CGG repeat expansions in neuronal intranuclear inclusion disease, oculopharyngodistal myopathy and an overlapping disease
Noncoding repeat expansions cause various neuromuscular diseases, including myotonic dystrophies, fragile X tremor/ataxia syndrome, some spinocerebellar ataxias, amyotrophic lateral sclerosis and benign adult familial myoclonic epilepsies. Inspired by the striking similarities in the clinical and neuroimaging findings between neuronal intranuclear inclusion disease (NIID) and fragile X tremor/ataxia syndrome caused by noncoding CGG repeat expansions in FMR1, we directly searched for repeat expansion mutations and identified noncoding CGG repeat expansions in NBPF19 (NOTCH2NLC) as the causative mutations for NIID. Further prompted by the similarities in the clinical and neuroimaging findings with NIID, we identified similar noncoding CGG repeat expansions in two other diseases: oculopharyngeal myopathy with leukoencephalopathy and oculopharyngodistal myopathy, in LOC642361/NUTM2B-AS1 and LRP12, respectively. These findings expand our knowledge of the clinical spectra of diseases caused by expansions of the same repeat motif, and further highlight how directly searching for expanded repeats can help identify mutations underlying diseases.