Summary: Washington State University researchers have developed a computer algorithm that can accurately map brain networks from electronic microscopy images.
Source: Washington State University.
A WSU research team for the first time has developed a computer algorithm that is nearly as accurate as people are at mapping brain neural networks — a breakthrough that could speed up the image analysis that researchers use to understand brain circuitry.
A report on the WSU team’s work currently in the journal, Bioinformatics.
Like mapping 100 billion homes
For more than a generation, people have been trying to improve understanding of human brain circuitry, but are challenged by its vast complexity. It is similar to having a satellite image of the earth and trying to map out 100 billion homes, all of the connecting streets and everyone’s destinations, said Shuiwang Ji, associate professor in the School of Electrical Engineering and Computer Science and lead researcher on the project.
Researchers, in fact, took more than a decade to fully map the circuitry of just one animal’s brain — a worm that has only 302 neurons. The human brain, meanwhile, has about 100 billion neurons, and the amount of data needed to fully understand its circuitry would require 1000 exabytes of data, or the equivalent of all the data that is currently available in the world.
Neuron by neuron
To map neurons, researchers currently use an electron microscope to take pictures — with one image usually containing a small number of neurons. The researchers then study each neuron’s shape and size as well as its thousands of connections with other nearby neurons to learn about its role in behavior or biology.
“We don’t know much about how brains work,” said Ji.
With such rudimentary understanding of our circuitry, researchers are limited in their ability to understand the causes of devastating brain diseases, such as Alzheimer’s, schizophrenia, autism or Parkinson’s disease, he said. Instead, they have to rely on trial and error experimentation to come up with treatments. The National Academy of Engineering has listed improving understanding of the human brain as one of its grand challenges for the 21st century.
Accurate as humans
In 2013, MIT organized a competition that called on researchers to develop automated computer algorithms that could speed up image analysis, decode and understand images of brain circuitry. As part of the competition, the algorithms are compared to work that was done by a real team of neuroscientists. If computers can become as accurate as humans, they will be able to do the computations much faster and cheaper than humans, said Ji.
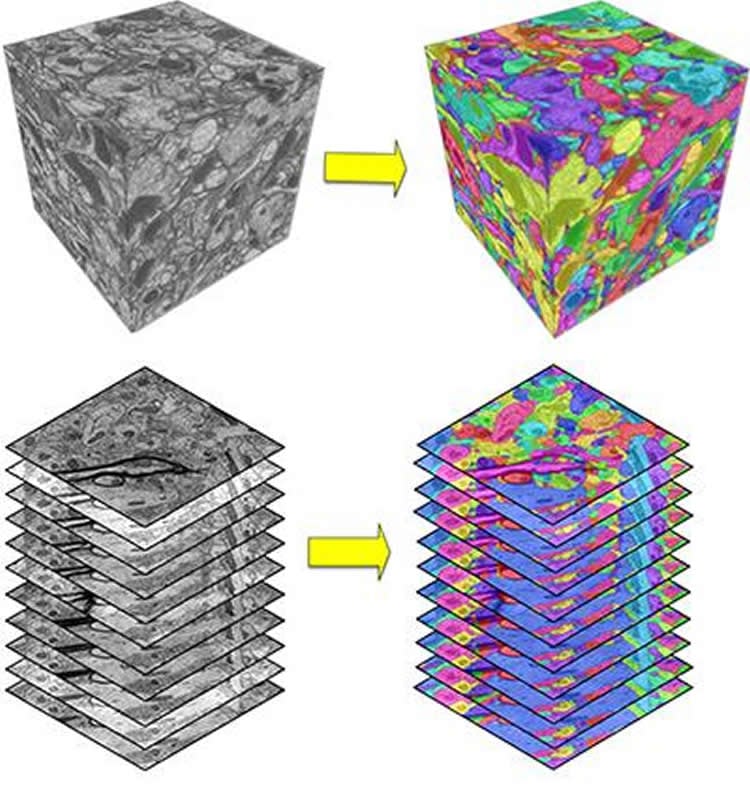
WSU’s research team developed the first computational model that was able to reach a human level of performance in accuracy.
Just as a human eye takes in information and then analyzes it in multiple stages, the WSU team developed a computational model that takes the image as its input and then processes it in a many-layered network before coming to a decision. In their algorithm, the researchers developed an artificial neural network that imitates humans’ complex biological neural networks.
While the WSU research team was able to approach human accuracy in the MIT challenge, they still have a lot of work to do in getting the computers to develop complete and accurate neural maps. The computers still make a large number of mistakes, and there is not yet a gold standard for comparing human and computational results, said Ji. Although it may not be realistic to expect that automated methods would completely replace human soon, improvements in computational methods will certainly lead to reduced manual proof-reading, he added.
Funding: The work was funded by Ji’s National Science Foundation CAREER award as well as from Washington State University. He also recently received a $400,000 National Institutes of Health grant to continue the work.
The work is in keeping with WSU’s Grand Challenges, a suite of research initiatives aimed at large societal issues. It is particularly relevant to the challenge of sustaining health and its theme of treating disease.
The code is available at https://github.com/divelab/deepem3d/
Source: Shuiwang Ji – Washington State University
Image Source: NeuroscienceNews.com image is credited to Washington State University.
Original Research: Abstract for “DeepEM3D: approaching human-level performance on 3D anisotropic EM image segmentation” by Tao Zeng, Bian Wu, and Shuiwang Ji in Bioinformatics. Published online August 15 2017 doi:10.1093/bioinformatics/btx188
[cbtabs][cbtab title=”MLA”]Washington State University”Computer Approaches Human Skill for First Time in Mapping Brain.” NeuroscienceNews. NeuroscienceNews, 17 August 2017.
<https://neurosciencenews.com/computer-skull-brain-mapping-7323/>.[/cbtab][cbtab title=”APA”]Washington State University(2017, August 17). Computer Approaches Human Skill for First Time in Mapping Brain. NeuroscienceNew. Retrieved August 17, 2017 from https://neurosciencenews.com/computer-skull-brain-mapping-7323/[/cbtab][cbtab title=”Chicago”]Washington State University”Computer Approaches Human Skill for First Time in Mapping Brain.” https://neurosciencenews.com/computer-skull-brain-mapping-7323/ (accessed August 17, 2017).[/cbtab][/cbtabs]
Abstract
DeepEM3D: approaching human-level performance on 3D anisotropic EM image segmentation
Motivation
Progress in 3D electron microscopy (EM) imaging has greatly facilitated neuroscience research in high-throughput data acquisition. Correspondingly, high-throughput automated image analysis methods are necessary to work on par with the speed of data being produced. One such example is the need for automated EM image segmentation for neurite reconstruction. However, the efficiency and reliability of current methods are still lagging far behind human performance.
Results
Here, we propose DeepEM3D, a deep learning method for segmenting 3D anisotropic brain electron microscopy images. In this method, the deep learning model can efficiently build feature representation and incorporate sufficient multi-scale contextual information. We propose employing a combination of novel boundary map generation methods with optimized model ensembles to address the inherent challenges of segmenting anisotropic images. We evaluated our method by participating in the 3D segmentation of neurites in EM images (SNEMI3D) challenge. Our submission is ranked #1 on the current leaderboard as of Oct 15, 2016. More importantly, our result was very close to human-level performance in terms of the challenge evaluation metric: namely, a Rand error of 0.06015 versus the human value of 0.05998.
“DeepEM3D: approaching human-level performance on 3D anisotropic EM image segmentation” by Tao Zeng, Bian Wu, and Shuiwang Ji in Bioinformatics. Published online August 15 2017 doi:10.1093/bioinformatics/btx188






