People with autism have a wide range of symptoms, with no two people sharing the exact type and severity of behaviors. Now a large-scale analysis of hundreds of patients and nearly 1000 genes has started to uncover how diversity among traits can be traced to differences in patients’ genetic mutations. The study, conducted by researchers at Columbia University Medical Center, was published Dec. 22 in the journal Nature Neuroscience.
Autism researchers have identified hundreds of genes that, when mutated, likely increase the risk of developing autism spectrum disorder (ASD). Much of the variability among people with ASD is thought to stem from the diversity of underlying genetic changes, including the specific genes mutated and the severity of the mutation.
“If we can understand how different mutations lead to different features of ASD, we may be able to use patients’ genetic profiles to develop accurate diagnostic and prognostic tools and perhaps personalize treatment,” said senior author Dennis Vitkup, PhD, associate professor of systems biology and biomedical informatics at Columbia University’s College of Physicians & Surgeons.
To investigate the links between genetic mutations and autism traits, Dr. Vitkup and a team of Columbia graduate students (Jonathan Chang, Sarah R. Gilman, and Andrew H. Chiang) analyzed genetic and clinical data on hundreds of patients with ASD from the Simons Simplex Collection.
IQ and gender differences in autism influenced by severity of mutations
The group found that more damaging genetic mutations usually lead to worse disease outcomes. “It looks as if high-IQ autism cases are usually triggered by milder mutations,” Dr. Vitkup said.
Patients with low-verbal or nonverbal IQs usually had mutations in genes that are more active in the brain. And high-IQ individuals were less likely to have mutations that completely shut down genes. Instead, mutations that only partially damage normal gene function in the brain appear to be predominantly associated with high-functioning autism cases.
Gender differences in autism could also be traced to the types of genes mutated in the individual. Though ASD is far more common in males, females with ASD are more likely to fall on the severe end of the spectrum.
The Columbia researchers found that the genes mutated in females generally had greater activity throughout the brain than those mutated in males. Very damaging ASD mutations in girls on average are found in genes that are almost twice as active as typical genes in normal brains.
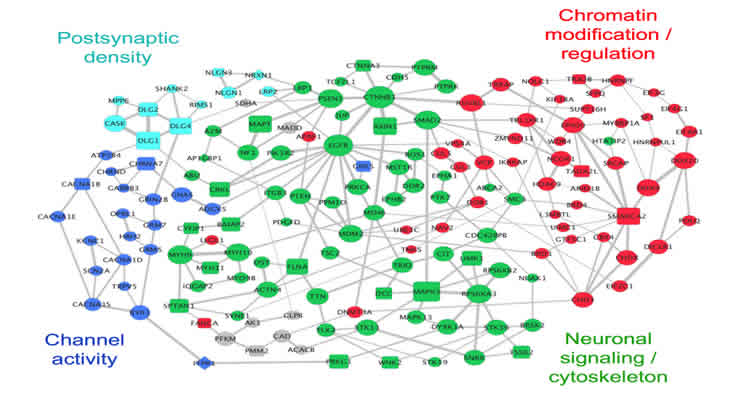
“These patterns are consistent with the idea that there are mechanisms that protect females,” Dr. Vitkup said. “Most often, only when a mutation hits a highly active gene do we see symptoms in females. Given that the inherent differences in gene activity in male and female brains are typically on the order of a few percent, these findings are quite remarkable.”
Autism mutations disrupt multiple cell types in brain
Behavioral variability in autism patients may also stem from the types of brain cells affected, and the Columbia researchers have taken the first steps in determining which cell types in the brain are most affected by autism mutations.
The team identified these cells by looking at the normal activity of autism-related genes in dozens of similar cell types in mouse brains. The analysis showed that many different types of neurons throughout the brain are affected by mutations in autism genes.
“The idea that eventually all autism mutations would converge onto a single type of neuron or single brain area isn’t what we see in the data,” Dr. Vitkup said. “Instead, an autism mutation usually affects multiple brain areas simultaneously.”
Certain neurons, however, appear to be more affected than others. The Columbia researchers found strong effects in cortical and striatal neurons that form a circuit that controls repetitive motions and behaviors, such as rocking, an insistence on sameness, and restricted interests, which are common in people with ASD.
“There are many hypotheses about the types of neurons and circuits involved in autism, but by using unbiased genome-wide approaches, like the one used in this study, one can understand which neurons are the most important and explain the core features we see in people with ASD,” said Dr. Vitkup.
Identifying the circuits involved is the next step in understanding autism, he said. “Huge progress has been made in the last five years: We and our colleagues have now identified multiple affected genes, and we are coming to a consensus about how the genes work together in biological networks. Now, based on the affected genes, we are identifying affected cell types and brains circuits and trying to connect them to disease outcomes in individual patients.”
The research was funded by the Simons Foundation (308962) and the NIH (U54CA121852 and T32GM082797).
The authors declare no financial or other conflicts of interest.
Contact: Lucky Tran – Columbia University Medical Center
Source: Columbia University Medical Center press release
Image Source: The image is credited to Dennis Vitkup and is adapted from the Columbia University Medical Center press release
Original Research: Abstract for “Genotype to phenotype relationships in autism spectrum disorders” by Jonathan Chang, Sarah R Gilman, Andrew H Chiang, Stephan J Sanders and Dennis Vitkup in Nature Neuroscience. Published online December 22 2014 doi:10.1038/nn.3907