Summary: A reconstituted circadian clock of cyanobacteria can run for consistent days, allowing researchers to study the interactions of the clock proteins in real-time and observe how the clock exerts control over gene expression.
Source: UC Santa Cruz
Daily cycles in virtually every aspect of our physiology are driven by biological clocks (also called circadian clocks) in our cells. The cyclical interactions of clock proteins keep the biological rhythms of life in tune with the daily cycle of night and day, and this happens not only in humans and other complex animals but even in simple, single-celled organisms such as cyanobacteria.
A team of scientists has now reconstituted the circadian clock of cyanobacteria in a test tube, enabling them to study rhythmic interactions of the clock proteins in real time and understand how these interactions enable the clock to exert control over gene expression.
Researchers in three labs at UC Santa Cruz, UC Merced, and UC San Diego collaborated on the study, published October 8 in Science.
“Reconstituting a complicated biological process like the circadian clock from the ground up has really helped us learn how the clock proteins work together and will enable a much deeper understanding of circadian rhythms,” said Carrie Partch, professor of chemistry and biochemistry at UC Santa Cruz and a corresponding author of the study.
Partch noted that the molecular details of circadian clocks are remarkably similar from cyanobacteria to humans. Having a functioning clock that can be studied in the test tube (“in vitro”) instead of in living cells (“in vivo”) provides a powerful platform for exploring the clock’s mechanisms and how it responds to changes. The team conducted experiments in living cells to confirm that their in vitro results are consistent with the way the clock operates in live cyanobacteria.
“These results were so surprising because it is common to have results in vitro that are somewhat inconsistent with what is observed in vivo. The interior of live cells is highly complex, in stark contrast to the much simpler conditions in vitro,” said Andy LiWang, professor of chemistry and biochemistry at UC Merced and a corresponding author of the paper.
The new study builds on previous work by Japanese researchers, who in 2005 reconstituted the cyanobacterial circadian oscillator, the basic 24-hour timekeeping loop of the clock. The oscillator consists of three related proteins: KaiA, KaiB, and KaiC. In living cells, signals from the oscillator are transmitted through other proteins to control the expression of genes in a circadian cycle.
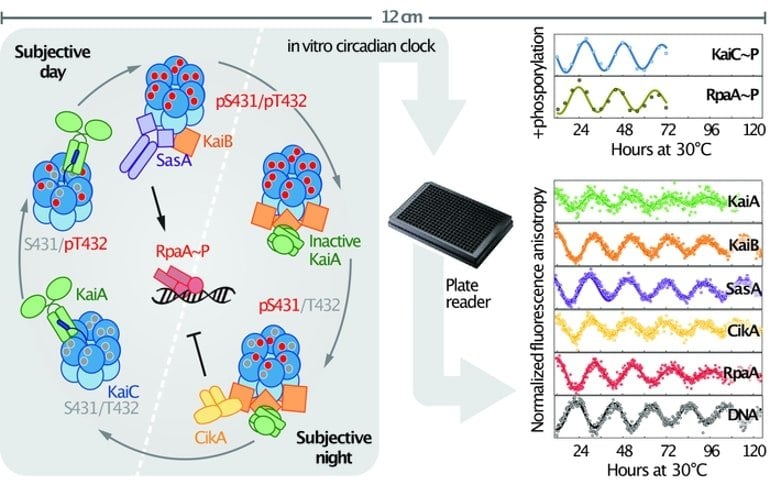
The new in vitro clock includes, in addition to the oscillator proteins, two kinase proteins (SasA and CikA), whose activities are modified by interacting with the oscillator, as well as a DNA-binding protein (RpaA) and its DNA target.
“SasA and CikA respectively activate and deactivate RpaA such that it rhythmically binds and unbinds DNA,” LiWang explained. “In cyanobacteria, this rhythmic binding and unbinding at over 100 different sites in their genome activates and deactivates the expression of numerous genes important to health and survival.”
Using fluorescent labeling techniques, the researchers were able to track the interactions between all of these clock components as the whole system oscillates with a circadian rhythm for many days and even weeks. This system enabled the team to determine how SasA and CikA enhance the robustness of the oscillator, keeping it ticking under conditions in which the KaiABC proteins by themselves would stop oscillating.
The researchers also used the in vitro system to explore the genetic origins of clock disruption in an arrhythmic strain of cyanobacteria. They identified a single mutation in the gene for RpaA that reduces the protein’s DNA-binding efficiency.
“A single amino acid change in the transcription factor makes the cell lose the rhythm of gene expression, even though its clock is intact,” said coauthor Susan Golden, director of the Center for Circadian Biology at UC San Diego, of which Partch and LiWang are also members.
“The real beauty of this project is how the team drawn from three UC campuses came together to pool approaches toward answering how a cell can tell time,” she added.
“The active collaboration extended well beyond the principal investigators, with the students and postdocs who were trained in different disciplines conferring among themselves to share genetics, structural biology, and biophysical data, explaining to one another the significance of their findings. The cross-discipline communication was as important to the success of the project as the impressive skills of the researchers.”
The authors of the paper include first authors Archana Chavan and Joel Heisler at UC Merced and Jeffrey Swan at UC Santa Cruz, as well as coauthors Cigdem Sancar, Dustin Ernst, and Mingxu Fang at UC San Diego, and Joseph Palacios, Rebecca Spangler, Clive Bagshaw, Sarvind Tripathi, and Priya Crosby at UC Santa Cruz.
Funding: This work was supported by the National Institutes of Health and the National Science Foundation.
About this circadian rhythm research news
Author: Tim Stephens
Source: UC Santa Cruz
Contact: Tim Stephens – UC Santa Cruz
Image: The image is credited to Andy LiWang
Original Research: The findings will appear in Science