Summary: New collaborative studies shed light on the organization of cells in key areas of the mouse brain and the organization of transcriptome, epigenomic, and regulatory factors that provide the brain cells with purpose and function.
Source: UCSD
The circuits of the human brain contain more than 100 billion neurons, each linked to many other neurons via thousands of synaptic connections, resulting in a three-pound organ that is profoundly more complex than the sum of its innumerable parts.
In recent years, however, transformative advances in imaging, sequencing and computational technologies have opened the possibility of mapping a human brain truly at the resolution of its molecular and cellular components. While that ultimate goal remains to be achieved, researchers have steadily progressed with a smaller, but no less momentous, effort: an atlas of the mouse brain.
In a special issue of Nature, publishing online October 7, 2021, researchers at the University of California San Diego, with colleagues across the country, describe their progress in collection of papers. Two of the papers, in which UC San Diego scientists served as senior authors, further refine the organization of cells within key regions of the mouse brain and, more critically, the organization of transcriptomic, epigenomic and regulatory factors and elements that provide these brain cells with function and purpose.
“To truly understand how the brain functions, and from that knowledge develop new drugs and therapies to improve human lives and health, we need to see and quantify brain structure, organization and function down to the level of single cells,” said Bing Ren, PhD, director of the Center for Epigenomics, professor of cellular and molecular medicine at UC San Diego School of Medicine and member of the Ludwig Institute for Cancer Research at UC San Diego.
“Depth and specificity are essential,” agreed Eran A. Mukamel, PhD, director of the Computational Neural DNA Dynamics Lab and associate professor in the Department of Cognitive Science at UC San Diego. “We want a comprehensive parts list for the brain, including not just the locations and connections of the neurons, but also the molecular and epigenetic fingerprints that give them their specialized identity.”
Gene regulatory elements
Since 2006, there has been a concerted, international effort to create a three-dimensional atlas of the mouse brain, which is roughly the size of a pea and comprised of approximately eight to 14 million neurons and glial cells. Though the mouse brain is not a miniature version of the human brain, it has proven to be a powerful model for studying many human brain functions, diseases and mental disorders, in part because the genes responsible for building and operating both human and rodent organs are 90 percent identical.
In their paper, senior author Ren, colleagues and collaborators at the Center for Epigenomics focused on creating an atlas of gene regulatory elements in the mouse cerebrum, the evolutionarily youngest region of the brain that supports high-level sensory perception, motor control and cognitive functions.
Recent surveys of mouse and human brains have revealed that the cerebrum contains hundreds of neural cell types distributed in different regions, but the transcriptional regulatory programs — the directions responsible for each cell’s unique pattern of gene expression, and hence its identity and function — remain unknown.
Ren’s team probed accessible chromatin — the stuff of chromosomes — in more than 800,000 individual cell nuclei from 45 locations in the adult mouse brain, then used the data to map the state of 491,818 candidate cis-regulatory DNA elements in 160 distinct cell types. Cis-regulatory elements are regions of non-coding DNA that regulate transcription (copying a segment of DNA into RNA) of neighboring genes.
They found that different types of neurons are located in distinct areas of the mouse brain, and the specificity of their spatial distribution and function is correlated, and likely driven, by the unique set of cis-regulatory DNA elements within each cell type. Indeed, some of the cell-type-specific elements identified by Ren’s team were independently shown to be sufficient to drive reporter gene expression in specific sub-classes of neurons in the mouse brain.
Surprisingly, most of the mouse brain cis-regulatory elements mapped by the researchers have homologous or similar sequences in the human genome that may act as regulatory elements, and therefore could be used to annotate gene regulatory elements involved in human brain cell type specification.
Ren said the findings provide a foundation for comprehensive analysis of gene regulatory programs of the mammalian brain, including humans, and can assist in interpreting noncoding risk variants that contribute to various neurological diseases and traits in humans.
Transcriptomic and epigenomic elements
Each cell or population of cells produces a unique pattern of RNA transcripts — strands of RNA transcribed from DNA that convey genetic instructions for the proteins that direct and sustain life. It’s estimated that millions of chemical reactions occur within mammalian cells every second. That complexity, combined with growing datasets describing the functions of genes, fats, proteins, sugars and other players in cell biology, have complicated efforts to understand how the brain is organized and functions.
Mukamel and colleagues brought together advanced sequencing techniques to focus on the mouse primary motor cortex, a brain region fundamental to movement. They generated more than 500,000 transcriptomes and epigenomes — comprehensive listings of all of the RNA molecules and modifications of DNA that make each mouse brain cell unique.
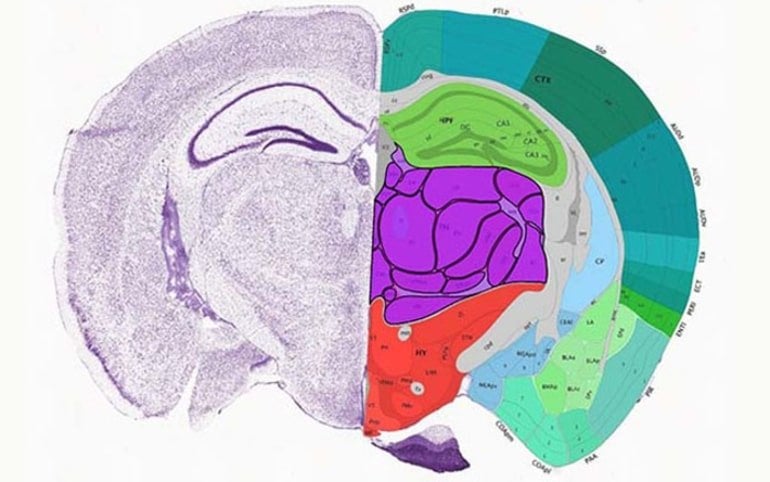
Using novel computational and statistical models, they created a multimodal atlas of 56 neuronal cell types in the mouse primary motor cortex that comprehensively describes their molecular, genomic and anatomic features.
Mukamel said the study showed that each brain cell has a coordinated pattern of gene expression and epigenetic regulation that can be recognized with high fidelity using different sequencing techniques. Just as an individual has characteristic handwriting, facial features, vocal patterns and personality traits, the authors found that the RNA and DNA signatures of cell types in the motor cortex differentiate each cell from its neighbors.
And just as our human individuality contributes to the strength and diversity of our communities, said Mukamel, the unique patterns of gene expression and regulation in brain circuits support a highly diverse network of cells with specialized roles and interdependent functions.
By combining both epigenomic and transcriptomic data from an unprecedented number of cells, Mukamel said the study demonstrates the potential of single-cell sequencing technologies to comprehensively map brain cell types — a lesson that will help in understanding the more complex circuits of the human brain.
About this brain mapping research news
Author: Scott La Fee
Source: UCSD
Contact: UCSD
Image: The image is credited to the Allen Brain Institute
Original Research: Open access.
“An atlas of gene regulatory elements in adult mouse cerebrum” by Bing Ren et al. Nature
Open access.
“A transcriptomic and epigenomic cell atlas of the mouse primary motor cortex” by Eran A. Mukamel et al. Nature
Abstract
An atlas of gene regulatory elements in adult mouse cerebrum
The mammalian cerebrum performs high-level sensory perception, motor control and cognitive functions through highly specialized cortical and subcortical structures.
Recent surveys of mouse and human brains with single-cell transcriptomics and high-throughput imaging technologies have uncovered hundreds of neural cell types distributed in different brain regions, but the transcriptional regulatory programs that are responsible for the unique identity and function of each cell type remain unknown.
Here we probe the accessible chromatin in more than 800,000 individual nuclei from 45 regions that span the adult mouse isocortex, olfactory bulb, hippocampus and cerebral nuclei, and use the resulting data to map the state of 491,818 candidate cis-regulatory DNA elements in 160 distinct cell types.
We find high specificity of spatial distribution for not only excitatory neurons, but also most classes of inhibitory neurons and a subset of glial cell types. We characterize the gene regulatory sequences associated with the regional specificity within these cell types.
We further link a considerable fraction of the cis-regulatory elements to putative target genes expressed in diverse cerebral cell types and predict transcriptional regulators that are involved in a broad spectrum of molecular and cellular pathways in different neuronal and glial cell populations.
Our results provide a foundation for comprehensive analysis of gene regulatory programs of the mammalian brain and assist in the interpretation of noncoding risk variants associated with various neurological diseases and traits in humans.
Abstract
A transcriptomic and epigenomic cell atlas of the mouse primary motor cortex
Single-cell transcriptomics can provide quantitative molecular signatures for large, unbiased samples of the diverse cell types in the brain. With the proliferation of multi-omics datasets, a major challenge is to validate and integrate results into a biological understanding of cell-type organization.
Here we generated transcriptomes and epigenomes from more than 500,000 individual cells in the mouse primary motor cortex, a structure that has an evolutionarily conserved role in locomotion. We developed computational and statistical methods to integrate multimodal data and quantitatively validate cell-type reproducibility.
The resulting reference atlas—containing over 56 neuronal cell types that are highly replicable across analysis methods, sequencing technologies and modalities—is a comprehensive molecular and genomic account of the diverse neuronal and non-neuronal cell types in the mouse primary motor cortex.
The atlas includes a population of excitatory neurons that resemble pyramidal cells in layer 4 in other cortical regions. We further discovered thousands of concordant marker genes and gene regulatory elements for these cell types.
Our results highlight the complex molecular regulation of cell types in the brain and will directly enable the design of reagents to target specific cell types in the mouse primary motor cortex for functional analysis.






