Summary: Researchers have succeeded in creating a new, bright red fluorescent protein which can be used to track essential cellular processes.
Source: University of Amsterdam.
After years of trying, biologists have succeeded in creating an extremely bright red fluorescent protein in the lab. This is good news for researchers, including cancer and stem cell researchers, who use fluorescent proteins to track essential cellular processes. The researchers at the University of Amsterdam, the Institut de Biologie structurale and the European Synchrotron in Grenoble describe their approach in the latest edition of the journal Nature Methods.
Biologists often employ a trick to understand how a (human) cell divides, secretes a hormone or transmits a signal to another cell. They attach small coloured lights to their proteins of interest in order to trace their movements and interactions in living cells under the microscope. The more colours of lights they use, the more processes they can track simultaneously.
Scientists used a fluorescent protein as a colour code in a cell for the first time in the 1990s. That protein was green and originated from a fluorescent jellyfish. By tinkering with that green protein, scientists were able to develop blue, turquoise and yellow variants in the years that followed. In the 2000s a red fluorescent protein was discovered in corals. However, nobody succeeded in developing a versatile and bright-red light in the cell – until now.
Protein mScarlet
Professor of Molecular Cytology Dorus Gadella and doctoral researchers Daphne Bindels and Lindsay Haarbosch have succeeded in creating a spectacularly bright-red fluorescent protein. They have dubbed the protein mScarlet and expect it to be used by research groups across the world, for example to gain a better understanding of how disruption of cellular processes causes uncontrolled cell division found in cancer cells. Gadella said ‘Just as other people study the stars and prepare future trips to Mars, we are exploring the universe of the proteins that regulate the cellular processes within our bodies.’
The research group created mScarlet by comparing the genetic blueprints of a range of red fluorescent proteins from corals. They searched for sequences that consistently occurred in the various genetic codes since these apparently are indispensable. The group assembled these essential pieces of code and then had a company synthesise a complete DNA strand. They introduced that synthetic DNA into a bacterium, which converted it into a protein.
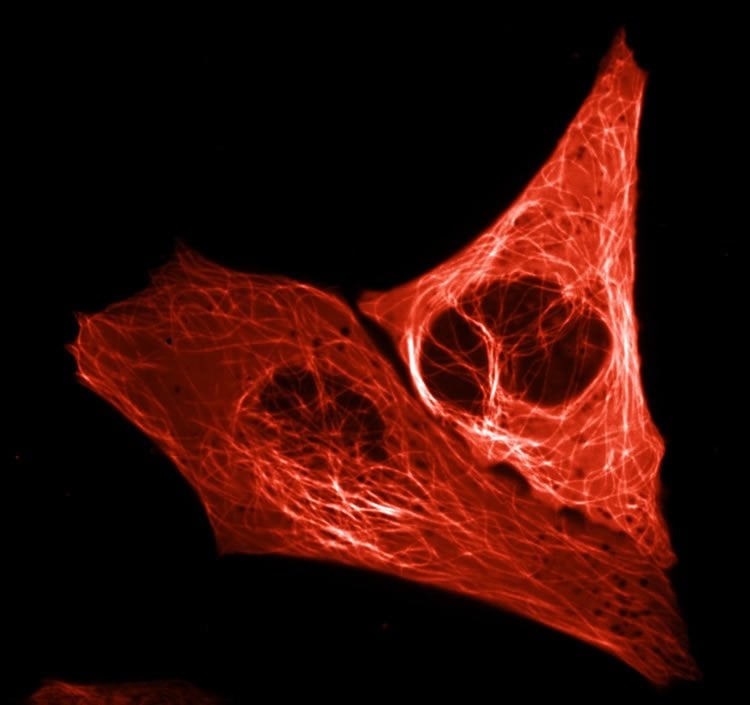
They assessed the brightness of each protein produced in this way under a microscope and then tinkered some more with the DNA code, observing how modifications affected the brightness. The entire process was a kind of lab-based evolutionary experiment which resulted in Gadella and his colleagues creating mScarlet, the protein with the highest brightness. That brightness will serve cellular microscopy well as it ensures the visibility of the proteins studied by scientists. Moreover, mScarlet is an ideal illuminating agent as it does not affect the functioning of the proteins to which it is tagged.
Cooperation with the Institut de Biologie Structurale and the European Synchrotron Radiation Facility
In order to fully understand mScarlet, the biologists eventually sent their red creation to the Institut de Biologie Structurale in Grenoble. A team of researchers led by structural biologist Antoine Royant utilized the European Synchrotron Radiation Facility (ESRF), one of the most powerful particle accelerators in the world, to reveal the molecular structure of the protein. Royant said ‘We discovered that mScarlet’s bright fluorescence is due to fact that the chromophore, the part of the molecule that absorbs light and then emits red light, is held rigidly flat by the protein wrapped around it.’
NeuroscienceNews.com would like to thank UvA Press Office for submitting this neuroscience research news to us for inclusion.
Source: UvA Press Office – University of Amsterdam
Image Source: NeuroscienceNews.com image is adapted from the University of Amsterdam press release.
Original Research: Abstract for “mScarlet: a bright monomeric red fluorescent protein for cellular imaging” by Daphne S Bindels, Lindsay Haarbosch, Laura van Weeren, Marten Postma, Katrin E Wiese, Marieke Mastop, Sylvain Aumonier, Guillaume Gotthard, Antoine Royant, Mark A Hink and Theodorus W J Gadella Jr in Nature Methods. Published online November 21 2016 doi:10.1038/nmeth.4074
[cbtabs][cbtab title=”MLA”]University of Amsterdam. “Bright Red Fluorescent Protein Created.” NeuroscienceNews. NeuroscienceNews, 23 November 2016.
<https://neurosciencenews.com/fluorescent-bright-red-protein-5588/>.[/cbtab][cbtab title=”APA”]University of Amsterdam. (2016, November 23). Bright Red Fluorescent Protein Created. NeuroscienceNews. Retrieved November 23, 2016 from https://neurosciencenews.com/fluorescent-bright-red-protein-5588/[/cbtab][cbtab title=”Chicago”]University of Amsterdam. “Bright Red Fluorescent Protein Created.” https://neurosciencenews.com/fluorescent-bright-red-protein-5588/ (accessed November 23, 2016).[/cbtab][/cbtabs]
Abstract
mScarlet: a bright monomeric red fluorescent protein for cellular imaging
We report the engineering of mScarlet, a truly monomeric red fluorescent protein with record brightness, quantum yield (70%) and fluorescence lifetime (3.9 ns). We developed mScarlet starting with a consensus synthetic template and using improved spectroscopic screening techniques; mScarlet’s crystal structure reveals a planar and rigidified chromophore. mScarlet outperforms existing red Förster proteins as a fusion tag, and it is especially useful as a fluorescence resonance energy transfer (FRET) acceptor in ratiometric imaging.
“mScarlet: a bright monomeric red fluorescent protein for cellular imaging” by Daphne S Bindels, Lindsay Haarbosch, Laura van Weeren, Marten Postma, Katrin E Wiese, Marieke Mastop, Sylvain Aumonier, Guillaume Gotthard, Antoine Royant, Mark A Hink and Theodorus W J Gadella Jr in Nature Methods. Published online November 21 2016 doi:10.1038/nmeth.4074