Summary: Study uncovers new genetic risk factors for age-related macular degeneration, a leading cause of vision loss in adults.
Source: PLOS
Combining a map of gene regulatory sites with disease-associated loci has uncovered a new genetic risk factor of adult-onset macular degeneration (AMD), according to a new study publishing January 17 in the open-access journal PLOS Biology by Ran Elkon and Ruth Ashery-Padan of Tel Aviv University, Israel, and colleagues.
The finding advances the understanding of the leading cause of visual impairment in adults.
AMD is caused by dysfunction in the retinal pigmented epithelium (RPE), a layer of tissue sandwiched between the photoreceptors that receive light, and the choriocapillaris, which nourishes the retina.
Because of the central importance of the RPE in AMD, the authors began by exploring a transcription factor (a protein that regulates specific genes) called LHX2 which, based on the team’s analysis of mouse mutants, is central to RPE development.
Knocking down LHX2 activity in RPE derived from human stem cells, they found that most affected genes were down-regulated, indicating that LHX2’s role was likely that of a transcriptional activator, binding to regulatory sites on the genome to increase activity of other genes.
The authors found that one affected gene, called OTX2, collaborated with LHX2 to regulate many genes in the RPE. By mapping the genomic sites that OTX2 and LHX2 could bind to, they showed that 68% of those that bound LHX2 were also bound by OTX2 (864 sites in all), suggesting they likely work together to promote the activity of a large suite of genes involved in RPE development and function.
A common method for finding genes that may contribute to a disease is to perform a genome-wide association study (GWAS), which identifies genome sequence differences between individuals (termed single nucleotide polymorphisms, or SNPs) that co-occur with disease.
Numerous such studies have previously been done in AMD. However, a GWAS by itself cannot uncover a causal mechanism.
Here, the authors compared their LHX2/OTX2 binding data to GWAS data in order to home in on variations that affected binding of the transcription factors, and thus may contribute to disease.
One such binding site was located within the promoter region of a gene called TRPM1, which had been previously linked to AMD, and found that the sequence variant at that site altered the binding strength of LHX2; the so-called C version bound it more strongly than the T version, and activity of the TRPM1 gene was higher when the C allele was present instead of the T allele.
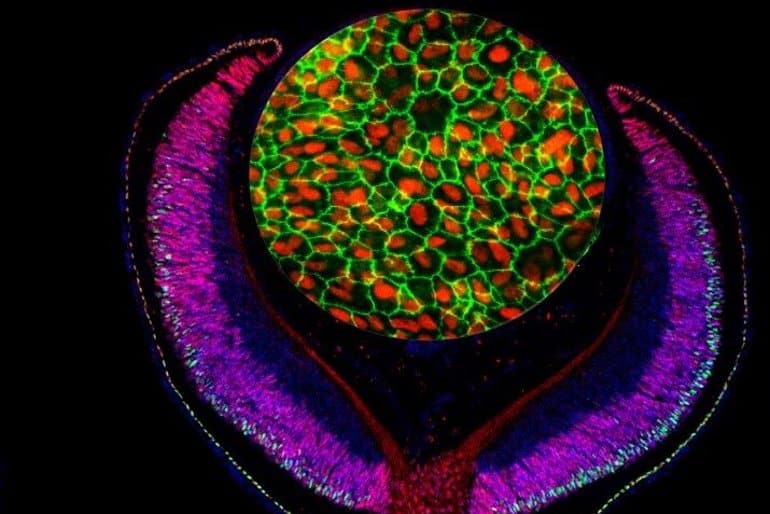
The results of the study indicate that the previously known increased risk of AMD from the variant identified in the GWAS was due to reduction in binding of the LHX2 transcription factor to the TRPM1 gene promoter, with a consequent reduction in activity of this gene.
The gene encodes a membrane ion channel, and previous studies have shown that mutations in the gene also cause visual impairment.
“Our study exemplifies how delineation of tissue-specific transcriptional regulators, their binding sites across the genome, and their downstream gene-regulatory networks can provide insights into a complex disease’s pathology,” the authors said.
Ashery-Padan adds, “The findings reveal a regulatory module consisting of LHX2 and OTX2 that controls the development and maintenance of the retinal pigmented epithelium, an important tissue of visual function.
“The genomic analyses further link the genomic regions bound by the two developmental factors to the genetics of the common, multifactorial blinding disease age-related macular degeneration (AMD).”
About this genetics and visual neuroscience research news
Author: Press Office
Source: PLOS
Contact: Press Office – PLOS
Image: The image is credited to Mazal Cohen-Gulkar, composite by Ruth Ashery-Padan
Original Research: Open access.
“The LHX2-OTX2 transcriptional regulatory module controls retinal pigmented epithelium differentiation and underlies genetic risk for age-related macular degeneration” by Ruth Ashery-Padan et al. PLOS Biology
Abstract
The LHX2-OTX2 transcriptional regulatory module controls retinal pigmented epithelium differentiation and underlies genetic risk for age-related macular degeneration
Tissue–specific transcription factors (TFs) control the transcriptome through an association with noncoding regulatory regions (cistromes). Identifying the combination of TFs that dictate specific cell fate, their specific cistromes and examining their involvement in complex human traits remain a major challenge.
Here, we focus on the retinal pigmented epithelium (RPE), an essential lineage for retinal development and function and the primary tissue affected in age-related macular degeneration (AMD), a leading cause of blindness.
By combining mechanistic findings in stem-cell-derived human RPE, in vivo functional studies in mice and global transcriptomic and proteomic analyses, we revealed that the key developmental TFs LHX2 and OTX2 function together in transcriptional module containing LDB1 and SWI/SNF (BAF) to regulate the RPE transcriptome.
Importantly, the intersection between the identified LHX2-OTX2 cistrome with published expression quantitative trait loci, ATAC-seq data from human RPE, and AMD genome-wide association study (GWAS) data, followed by functional validation using a reporter assay, revealed a causal genetic variant that affects AMD risk by altering TRPM1 expression in the RPE through modulation of LHX2 transcriptional activity on its promoter.
Taken together, the reported cistrome of LHX2 and OTX2, the identified downstream genes and interacting co-factors reveal the RPE transcription module and uncover a causal regulatory risk single-nucleotide polymorphism (SNP) in the multifactorial common blinding disease AMD.