In their struggle to survive and prosper, multicellular organisms rely on a complex network of communication between cells, which in humans are believed to number about 40 trillion. Now, in a study published in Nature Communications, a research group led by scientists from the RIKEN Center for Life Science Technologies (CLST) has published an overall map of how the cells in the human body communicate by systematically analyzing the relationship between ligands—substances such as insulin and interferon that embody messages between cells, and receptors—the proteins on cell surfaces that receive these messages when bound by the ligands.
The group looked at the expression of 1,894 ligand-receptor pairs—based on 642 ligands and 589 receptors—that have been reported in the literature so far, and drew up a large-scale map of cell-to-cell communication between 144 primary human cell types. The survey found that most cells express tens or even hundreds of ligands and receptors, creating a highly connected signaling network made up of cell types that can communicate with each other through multiple ligand-receptor paths.
The development of multicellular organisms from unicellular ancestors is one of the most profound evolutionary events in the history of life on Earth. During this transition, the cells of multicellular organisms began to communicate with each another in many ways, playing specific roles in the development and then control of their cellular functions. This process is particularly critical during early embryonic development when a cell’s differentiation and ultimate fate are controlled by communication with neighboring cells. In the developed organism, intercellular communication coordinates the activities between multiple cell types and makes organism-wide processes such as immune response, growth, and homeostasis. Defects in cell-cell communication, including the dysregulation of autocrine signaling, are also implied in the development of cancer, and autoimmune and metabolic diseases.
Despite the importance of this process, studies in intercellular communication between specialized cells in higher organisms such as mammals have generally focused on communication between just a few cell types and have been limited in scope to small numbers of ligand-receptor pairs. Currently there are no reports based on systematic studies attempting to elucidate and quantify the repertoire of signaling routes between different cell types.
To address this, the group looked at gene-expression data measured by the CAGE method in the RIKEN-led FANTOM5 project, using information from existing databases and large numbers of past publications, to generate the first large-scale draft map of primary cell-to-cell interactions.
Based on the analysis, the authors gained new insights into how cells communicate. According to Jordan Ramilowski, the first author of the study, “One intriguing conclusion is that signals between cells of the same type are surprisingly common, accounting for approximately two-thirds of cell-cell partners. We also discovered that receptors in general seem to have evolved earlier than ligands.”
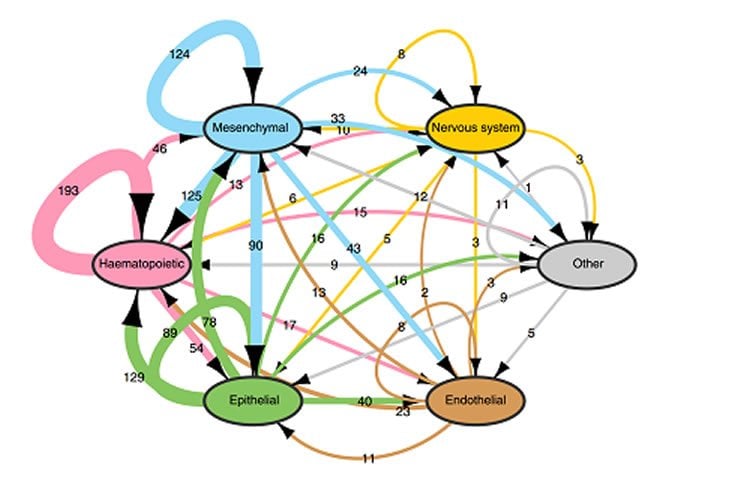
According to Alistair Forrest, who led the project, “Elucidating how cells communicate in an organism can contribute to the development of medical treatments. In particular, this data can be a key to discovering receptors that can be targets for therapies in various diseases.”
Considering the importance of these findings for the development of new medical therapeutics, the team has decided to publish the data online.
Source: Jens Wilkinson – RIKEN
Image Credit: The image is credited to the researchers
Original Research: Full open access research for “A draft network of ligand–receptor-mediated multicellular signalling in human” by Ramilowski J, Goldberg T, Harshbarger J, Kloppman E, Lizio M, Satagopam V, Itoh M, Kawaji H, Carninci P, Rost B, and Forrest A. in Nature Communications. Published online July 22 2015 doi:10.1038/ncomms8866
Abstract
A draft network of ligand–receptor-mediated multicellular signalling in human
Cell-to-cell communication across multiple cell types and tissues strictly governs proper functioning of metazoans and extensively relies on interactions between secreted ligands and cell-surface receptors. Herein, we present the first large-scale map of cell-to-cell communication between 144 human primary cell types. We reveal that most cells express tens to hundreds of ligands and receptors to create a highly connected signalling network through multiple ligand–receptor paths. We also observe extensive autocrine signalling with approximately two-thirds of partners possibly interacting on the same cell type. We find that plasma membrane and secreted proteins have the highest cell-type specificity, they are evolutionarily younger than intracellular proteins, and that most receptors had evolved before their ligands. We provide an online tool to interactively query and visualize our networks and demonstrate how this tool can reveal novel cell-to-cell interactions with the prediction that mast cells signal to monoblastic lineages via the CSF1–CSF1R interacting pair.
“A draft network of ligand–receptor-mediated multicellular signalling in human” by Ramilowski J, Goldberg T, Harshbarger J, Kloppman E, Lizio M, Satagopam V, Itoh M, Kawaji H, Carninci P, Rost B, and Forrest A. in Nature Communications. Published online July 22 2015 doi:10.1038/ncomms8866